Algal Genomes
Several nuclear genome sequencing projects have been completed to date, including those for:
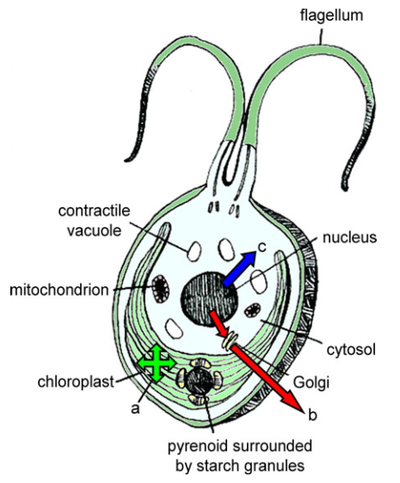
Chlamydomonas reinhardtii - one of the most commonly used eukaryotic model alga, a ~10-μm, unicellular, soil-dwelling green alga with multiple mitochondria, two anterior flagella for motility and mating, and a chloroplast that houses the photosynthetic apparatus and critical metabolic pathways. All three genomes (chloroplast, mitochondrial, and nuclear) can be transformed in Chlamydomonas, and each has distinct transcriptional, translational, and post-translational properties that make them distinct. Each of these genomes has been fully sequenced, providing a wealth of information and a strong foundation for targeted manipulation (Maul et al. 2002; Popescu and Lee 2007; Merchant et al. 2007).
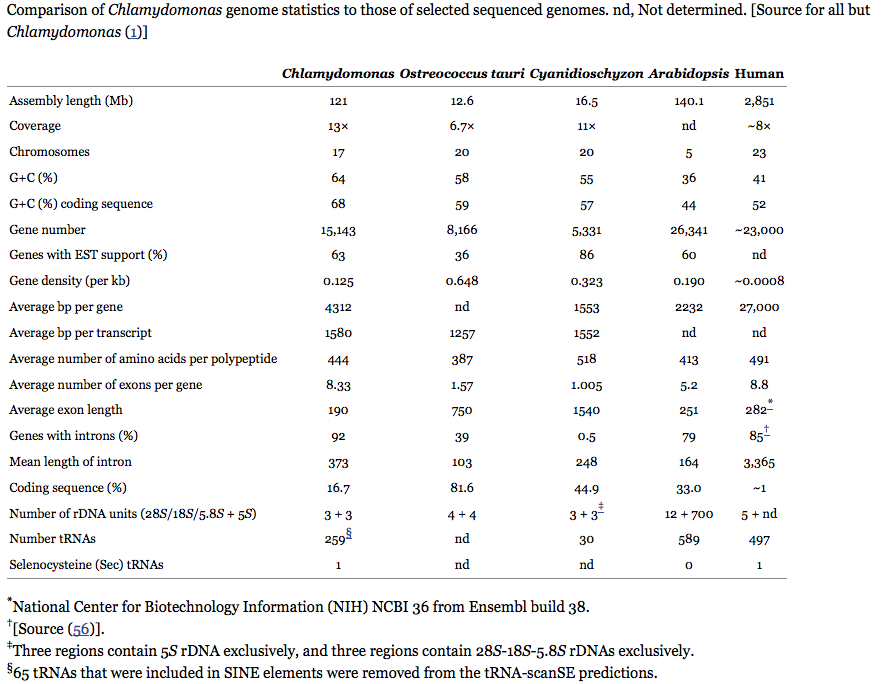
- Nucleus (17 linear scaffolds ~ 120 megabases (Mb), GC content of coding sequence = 64%, 14488 genes) - Merchant, S. S., S. E. Prochnik, O. Vallon, E. H. Harris, S. J. Karpowicz, G. B. Witman, A. Terry, A. Salamov, L. K. Fritz-Laylin, L. Marechal- Drouard, W. F. Marshall, L. H. Qu, D. R. Nelson, A. A. Sanderfoot, M. H. Spalding, V. V. Kapitonov, Q. Ren, P. Ferris, E. Lindquist, H. Shapiro, S. M. Lucas, J. Grimwood, J. Schmutz, P. Cardol, H. Cerutti, G. Chanf- reau, C. L. Chen, V. Cognat, M. T. Croft, R. Dent., S. Dutcher, E. Fernan- dez, H. Fukuzawa, D. Gonzalez-Ballester, D. Gonzalez-Halphen, A. Hall- mann, M. Hanikenne, M. Hippler, W. Inwood, K. Jabbari, M. Kalanon, R. Kuras, P. A. Lefebvre, S. D. Lemaire, A. V. Lobanov, M. Lohr, A. Manuell, I. Meier, L. Mets, M. Mittag, T. Mittelmeier, J. V. Moroney, J. Moseley, C. Napoli, A. M. Nedelcu, K. Niyogi, S. V. Novoselov, I. T. Paulsen, G. Pazour, S. Purton, J. P. Ral, D. M. Riano-Pachon, W. Riekhof, L. Rymarquis, M. Schroda, D. Stern, J. Umen, R. Willows, N. Wilson, S. L. Zimmer, J. Allmer, J. Balk, K. Bisova, C. J. Chen, M. Elias, K. Gendler, C. Hauser, M. R. Lamb, H. Ledford, J. C. Long, J. Minagawa, M. D. Page, J. Pan, W. Pootakham, S. Roje, A. Rose, E. Stahlberg, A. M. Terauchi, P. Yang, S. Ball, C. Bowler, C. L. Dieckmann, V. N. Gladyshev, P. Green, R. Jorgensen, S. Mayfield, B. Mueller-Roeber, S. Rajamani, R. T. Sayre, P. Brokstein, I. Dubchak, D. Goodstein, L. Hornick, Y. W. Huang, J. Jhaveri, Y. Luo, D. Martinez, W. C. Ngau, B. Otillar, A. Poliakov, A. Porter, L. Szajkowski, G. Werner, K. Zhou, I. V. Grigoriev, D. S. Rokhsar, and A. R. Grossman. 2007. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318:245–250.
- Mitochondria (Linear - 15758 bp, GC content of coding sequence = 21%, 1450 genes) - Popescu, C. E., & Lee, R. W. (2007). Mitochondrial genome sequence evolution in Chlamydomonas. Genetics, 175(2), 819–26. doi:10.1534/genetics.106.063156
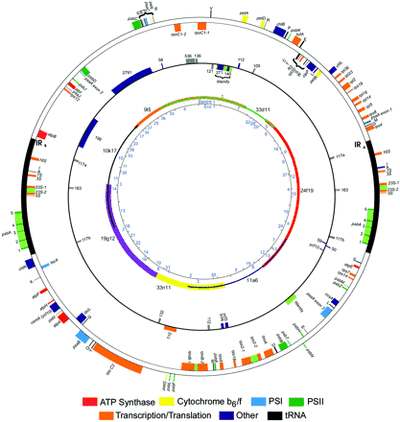
- Chloroplast/Plastid (Circular - 203828 bp, GC content of coding sequence = 34.6%, 102 genes, 99 expressed sequences (not including genes duplicated in the inverted repeat). This is the smallest plastid gene repertoire reported in the photosynthetic Viridiplantae and contrasts with 127 chloroplast genes in the green alga N. olivacea (Turmel et al., 1999) and 251 in the most gene-rich plastid DNA, that of the red alga Porphyra purpurea (Reith and Munholland, 1995). The G+C content is comparable to that of Chlorella (31.6%) and Arabidopsis (36.3%). The genome possesses two copies of an inverted repeat sequence (22,211 bp) - Maul, J. E., Lilly, J. W., Cui, L., Claude, W., Miller, W., Harris, E. H., & Stern, D. B. (2002). The Chlamydomonas reinhardtii Plastid Chromosome : Islands of Genes in a Sea of Repeats. The Plant Cell, 14(November), 2659–2679. doi:10.1105/tpc.006155.present
- See also: Shrager, J., C. Hauser, C. W. Chang, E. H. Harris, J. Davies, J. McDermott, R. Tamse, Z. Zhang, and A. R. Grossman. 2003. Chlamydomonas reinhardtii genome project. A guide to the generation and use of the cDNA information. Plant Physiol. 131:401–408
Phaeodactylum tricornutum - has the unusual property of being pleiomorphic (can occur in different shape and form), and this plasticity is related to the atypical nature of the cell wall, which is only poorly silicified compared with other diatoms (Lewin et al. 1958, Borowitzka and Volcani 1978). P. tricornutum exists in three different morphological forms: oval, fusiform, and triradiate. Basic biological features of P. tricornutum are relatively poorly studied, in particular its life cycle and its ecological relevance. The species appears to be unique in that it does not have an obligate requirement for silicic acid. The 27.4 Mb P. tricornutum nuclear genome currently exists as 33 individual scaffolds. These scaffolds are described as chromosomes in reports on the genome sequence, although ongoing work to completely close the genome sequence may collapse some of these scaffolds without telomere-to-telomere coverage into single chromosomes.
- Nucleus (33 linear scaffolds ~ 27.45 Mb, GC content of coding sequence = 48.8%, 10398 genes) Bowler, C., A. E. Allen, J. H. Badger, J. Grimwood, K. Jabbari, A. Kuo, U. Maheswari, C. Martens, F. Maumus, R. P. Otillar, E. Rayko, A. Salamov, K. Vandepoele, B. Beszteri, A. Gruber, M. Heijde, M. Katinka, T. Mock, K. Valentin, F. Verret, J. A. Berges, C. Brownlee, J. P. Cadoret, A. Chiovitti, C. J. Choi, S. Coesel, A. De Martino, J. C. Detter, C. Durkin, A. Falciatore, J. Fournet, M. Haruta, M. J. Huysman, B. D. Jenkins, K. Jiroutova, R. E. Jorgensen, Y. Joubert, A. Kaplan, N. Kroger, P. G. Kroth, J. La Roche, E. Lindquist, M. Lommer, V. Martin-Jezequel, P. J. Lopez, S. Lucas, M. Mangogna, K. McGinnis, L. K. Medlin, A. Montsant, M. P. Oudot-Le Secq, C. Napoli, M. Obornik, M. S. Parker, J. L. Petit, B. M. Porcel, N. Poulsen, M. Robison, L. Rychlewski, T. A. Rynearson, J. Schmutz, H. Shapiro, M. Siaut, M. Stanley, M. R. Sussman, A. R. Taylor, A. Vardi, P. von Dassow, W. Vyverman, A. Willis, L. S. Wyrwicz, D. S. Rokhsar, J. Weissenbach, E. V. Armbrust, B. R. Green, Y. Van de Peer, and I. V. Grigoriev. 2008. The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456:239–244.)
- Mitochondria (Linear - 77356 bp, GC content of coding sequence = 35%, 61 genes)
- Chloroplast/Plastid (Circular - 117369 bp, GC content of coding sequence = 32.6%, 170 genes)
Thalassiosira pseudonana (Armbrust, E. V., J. A. Berges, C. Bowler, B. R. Green, D. Martinez, N. H. Putnam, S. Zhou, A. E. Allen, K. E. Apt, M. Bechner, M. A. Brzezinski, B. K. Chaal, A. Chiovitti, A. K. Davis, M. S. Demarest, J. C. Detter, T. Glavina, D. Goodstein, M. Z. Hadi, U. Hellsten, M. Hildebrand, B. D. Jenkins, J. Jurka, V. V. Kapitonov, N. Kroger, W. W. Lau, T. W. Lane, F. W. Larimer, J. C. Lippmeier, S. Lucas, M. Medina, A. Montsant, M. Obornik, M. S. Parker, B. Palenik, G. J. Pazour, P. M. Richardson, T. A. Rynearson, M. A. Saito, D. C. Schwartz, K. Thamatrakoln, K. Valentin, A. Vardi, F. P. Wilkerson, and D. S. Rokhsar. 2004. The genome of the diatom Thalassio- sira pseudonana: ecology, evolution, and metabolism. Science 306:79–86)
Cyanidioschyzon merolae (Matsuzaki, M., O. Misumi, T. Shin-i, S. Maruyama, M. Takahara, S.-Y. Miyagishima, T. Mori, K. Nishida, F. Yagisawa, K. Nishida, Y. Yoshida, Y. Nishimura, S. Nakao, T. Kobayashi, Y. Momoyama, T. Higashiyama, A. Minoda, M. Sano, H. Nomoto, K. Oishi, H. Hayashi, F. Ohta, S. Nishizaka, S. Haga, S. Miura, T. Morishita, Y. Kabeya, K. Terasawa, Y. Suzuki, Y. Ishii, S. Asakawa, H. Takano, N. Ohta, H. Kuroiwa, K. Tanaka, N. Shimizu, S. Sugano, N. Sato, H. Nozaki, N. Ogasawara, Y. Kohara, and T. Kuroiwa. 2004. Genome sequence of the ultrasmall unicellular red alga Cyanidios- chyzon merolae 10D. Nature 428:653–657.)
Ostreococcus lucimarinus (Palenik, B., J. Grimwood, A. Aerts, P. Rouze, A. Salamov, N. Putnam, C. Dupont, R. Jorgensen, E. Derelle, S. Rombauts, K. Zhou, R. Otillar, S. S. Merchant, S. Podell, T. Gaasterland, C. Napoli, K. Gendler, A. Manuell, V. Tai, O. Vallon, G. Piganeau, S. Jancek, M. Heijde, K. Jabbari, C. Bowler, M. Lohr, S. Robbens, G. Werner, I. Dubchak, G. J. Pazour, Q. Ren, I. Paulsen, C. Delwiche, J. Schmutz, D. Rokhsar, Y. Van de Peer, H. Moreau, and I. V. Grigoriev. 2007. The tiny eukaryote Ostreococcus provides genomic insights into the paradox of plankton speciation. Proc. Natl. Acad. Sci. U. S. A. 104:7705–7710.)
Ostreococcus tauri (Derelle, E., C. Ferraz, S. Rombauts, P. Rouze, A. Z. Worden, S. Robbens, F. Partensky, S. Degroeve, S. Echeynie, R. Cooke, Y. Saeys, J. Wuyts, K. Jabbari, C. Bowler, O. Panaud, B. Piegu, S. G. Ball, J. P. Ral, F. Y. Bouget, G. Piganeau, B. De Baets, A. Picard, M. Delseny, J. Demaille, Y. Van de Peer, and H. Moreau. 2006. Genome analysis of the smallest free-living eukaryote Ostreococcus tauri unveils many unique features. Proc. Natl. Acad. Sci. U. S. A. 103:11647–11652.)
Ostreococcus tauri (Palenik, B., Grimwood, J., Aerts, A., Rouzé, P., Salamov, A., Putnam, N., … Grigoriev, I. V. (2007). The tiny eukaryote Ostreococcus provides genomic insights into the paradox of plankton speciation. Proceedings of the National Academy of Sciences of the United States of America, 104(18), 7705–10. doi:10.1073/pnas.0611046104)
Micromonas pusilla (Worden, A. Z., J. H. Lee, T. Mock, P. Rouze, M. P. Simmons, A. L. Aerts, A. E. Allen, M. L. Cuvelier, E. Derelle, M. V. Everett, E. Foulon, J. Grim- wood, H. Gundlach, B. Henrissat, C. Napoli, S. M. McDonald, M. S. Parker, S. Rombauts, A. Salamov, P. Von Dassow, J. H. Badger, P. M. Coutinho, E. Demir, I. Dubchak, C. Gentemann, W. Eikrem, J. E. Gready, U. John, W. Lanier, E. A. Lindquist, S. Lucas, K. F. Mayer, H. Moreau, F. Not, R. Otillar, O. Panaud, J. Pangilinan, I. Paulsen, B. Piegu, A. Poliakov, S. Robbens, J. Schmutz, E. Toulza, T. Wyss, A. Zelensky, K. Zhou, E. V. Armbrust, D. Bhattacharya, U. W. Goodenough, Y. Van de Peer, and I. V. Grigoriev. 2009. Green evolution and dynamic adaptations revealed by genomes of the marine picoeukaryotes Micromonas. Science 324:268–272.).
Ongoing microalgal genome sequencing projects include those for:
Fragilariopsis cylindrus
Pseudonitzschia
Thalassiosira rotula
Botryococcus braunii
Chlorella vulgaris
Dunaliella salina
Micromonas pusilla
Galdieria sulphuraria
Porphyra purpurea
Volvox carteri
Aureococcus anophageferrens
Source: Liolios, K., K. Mavromatis, N. Tavernarakis, and N. C. Kyrpides. 2008. The Genomes On Line Database (GOLD) in 2007: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res. 36:D475–D479.
Interesting observations have been made after genome sequencing microalgae: Diatoms such as Phaeodactylum tricornutum (affectionately referred to as "P.T." by researchers) are as far as 55% divergent from the centric diatom Thalassiosira pseudonana (affectionately referred to as "T.P." by researchers) genetically - that is almost as divergent as Phaeodactylum tricornutum is to a common water mold (Phytophthora sojae) - Table 2.
To put that into perspective these 2 diatoms are as divergent from each other as humans (Homo sapiens) are in comparison to a sea squirt (Ciona intestinalis) or a pufferfish (Takifugu rubripes):
To put that into perspective these 2 diatoms are as divergent from each other as humans (Homo sapiens) are in comparison to a sea squirt (Ciona intestinalis) or a pufferfish (Takifugu rubripes):
In addition, there are several completed and ongoing efforts to sequence plastid and mitochondrial genomes, as well as dynamic transcriptomes from many different microalgae (see 4, 9, 29, 66, 73, 101, 105, 130, 133, 149, 161, 169, 171, 196, 198 - references from Radakovits 2010 - Eukaryotic cell).
See also: http://www.jcvi.org/cms/publications/listing/abstract/article/the-comprehensive-microbial-resource/
See also: http://www.jcvi.org/cms/publications/listing/abstract/article/the-comprehensive-microbial-resource/