Genetic modifications in prokaryotic bacteria:
Cyanobacteria constitute one of the largest groups of bacteria with many hundreds of strains deposited in culture collections worldwide (Cohen and Gurevitz, 2006). The methods available for gene transfer to cyanobacteria are natural transformation, chemical transformation, electroporation, and conjugation. Among the many strains that are genetically amenable in this manner (Koksharova and Wolk, 2002) only a few strains dominate the research on genetic and metabolic manipulation (Flores et al., 2008; Heidorn et al., 2011). These strains include the unicellular, freshwater strains Synechocystis sp. PCC 6803 (Ikeuchi and Tabata, 2001) and Synechococcus elongatus sp. PCC 7942 (Holtman et al., 2005) that both are naturally transformable. Synechocystis sp. PCC 6803 was the first photosynthetic organism to have its genome sequenced and is one of the best characterized cyanobacteria. A commercial kit for engineering expression of recombinant proteins specifically in S. elongatus sp. PCC 7942 is now available (Thermo Fisher Scientific, previously Life Technologies Corp, Carlsbad, CA). The physiology and genome sequence of another model organism, S. elongatus sp. PCC 6301 is nearly identical to that of S. elongatus sp. PCC 7942.
The unicellular, marine strains Synechococcus sp. PCC 7002 (Frigaard et al., 2004) and Thermosynechococcus elongatus BP-1 (Iwai et al., 2004) are also naturally transformable although the latter strain has a higher transformation efficiency by electroporation. T. elongatus BP-1 is the only thermophilic strain (growth optimum 55°C) commonly used in genetic and physiological studies of cyanobacteria. Anabaena variabilis ATCC 29413 (Happe et al., 2000), Anabaena (also called Nostoc) sp. PCC 7120 (Kaneko et al., 2001), and Nostoc punctiforme ATCC 29133 (also called PCC 73102) (Meeks et al., 2001), are all filamentous, freshwater strains that perform N2 fixation in specialized cells called heterocysts. Gene transfer to these three strains is typically performed by conjugation.
With respect to physiology, cyanobacteria have certain advantages over algae and plants: cyanobacteria exhibit rapid growth and have less complex intracellular structures and cell walls. Cyanobacteria have a bacterial-type peptidoglycan cell wall unlike the complex carbohydrate cell wall found in algae and plants. Finally, some cyanobacteria are capable of using N2 as the sole source of nitrogen–something that algae and plants cannot (unless they have symbiotic N2-fixing bacteria).
Source: Rosgaard, L., de Porcellinis, A. J., Jacobsen, J. H., Frigaard, N.-U., & Sakuragi, Y. (2012). Bioengineering of carbon fixation, biofuels, and biochemicals in cyanobacteria and plants. Journal of Biotechnology, 162(1), 134–47. doi:10.1016/j.jbiotec.2012.05.006
Cyanobacteria constitute one of the largest groups of bacteria with many hundreds of strains deposited in culture collections worldwide (Cohen and Gurevitz, 2006). The methods available for gene transfer to cyanobacteria are natural transformation, chemical transformation, electroporation, and conjugation. Among the many strains that are genetically amenable in this manner (Koksharova and Wolk, 2002) only a few strains dominate the research on genetic and metabolic manipulation (Flores et al., 2008; Heidorn et al., 2011). These strains include the unicellular, freshwater strains Synechocystis sp. PCC 6803 (Ikeuchi and Tabata, 2001) and Synechococcus elongatus sp. PCC 7942 (Holtman et al., 2005) that both are naturally transformable. Synechocystis sp. PCC 6803 was the first photosynthetic organism to have its genome sequenced and is one of the best characterized cyanobacteria. A commercial kit for engineering expression of recombinant proteins specifically in S. elongatus sp. PCC 7942 is now available (Thermo Fisher Scientific, previously Life Technologies Corp, Carlsbad, CA). The physiology and genome sequence of another model organism, S. elongatus sp. PCC 6301 is nearly identical to that of S. elongatus sp. PCC 7942.
The unicellular, marine strains Synechococcus sp. PCC 7002 (Frigaard et al., 2004) and Thermosynechococcus elongatus BP-1 (Iwai et al., 2004) are also naturally transformable although the latter strain has a higher transformation efficiency by electroporation. T. elongatus BP-1 is the only thermophilic strain (growth optimum 55°C) commonly used in genetic and physiological studies of cyanobacteria. Anabaena variabilis ATCC 29413 (Happe et al., 2000), Anabaena (also called Nostoc) sp. PCC 7120 (Kaneko et al., 2001), and Nostoc punctiforme ATCC 29133 (also called PCC 73102) (Meeks et al., 2001), are all filamentous, freshwater strains that perform N2 fixation in specialized cells called heterocysts. Gene transfer to these three strains is typically performed by conjugation.
With respect to physiology, cyanobacteria have certain advantages over algae and plants: cyanobacteria exhibit rapid growth and have less complex intracellular structures and cell walls. Cyanobacteria have a bacterial-type peptidoglycan cell wall unlike the complex carbohydrate cell wall found in algae and plants. Finally, some cyanobacteria are capable of using N2 as the sole source of nitrogen–something that algae and plants cannot (unless they have symbiotic N2-fixing bacteria).
Source: Rosgaard, L., de Porcellinis, A. J., Jacobsen, J. H., Frigaard, N.-U., & Sakuragi, Y. (2012). Bioengineering of carbon fixation, biofuels, and biochemicals in cyanobacteria and plants. Journal of Biotechnology, 162(1), 134–47. doi:10.1016/j.jbiotec.2012.05.006
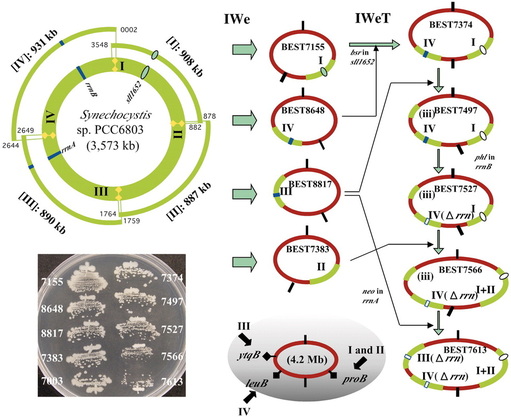
In 2005 Itaya and coworkers developed a method for cloning megabase-sized segments of DNA into the Bacillus subtilis genome using the natural transformation system of this bacterium. Almost all of the cyanobacterial Synechocystis PCC6803 genome was cloned as a set of four separate ~800-kb to 900-kb fragments into the B. subtilis chromosome by a reiterated “inch worm” process to generate a composite genome.
Source: M. Itaya, K. Tsuge, M. Koizumi, K. Fujita, Proc. Natl. Acad. Sci. U.S.A. 102, 15971 (2005).
Source: M. Itaya, K. Tsuge, M. Koizumi, K. Fujita, Proc. Natl. Acad. Sci. U.S.A. 102, 15971 (2005).
Transfer of genomes from photosynthetic organisms into yeast for precise genome modifications:
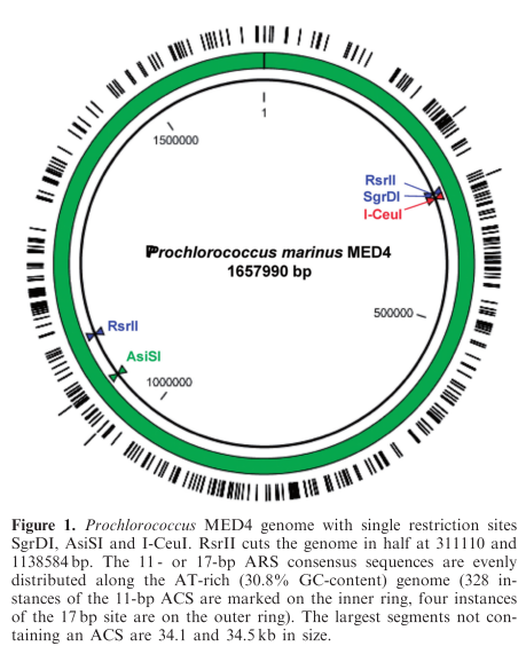
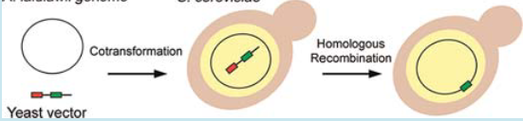
Marine cyanobacteria of the genus Prochlorococcus represent numerically dominant photoautotrophs residing throughout the euphotic zones in the open oceans and are major contributors to the global carbon cycle. Prochlorococcus has remained a genetically intractable bacterium due to slow growth rates and low transformation efficiencies using standard techniques. Our recent successes in cloning and genetically engineering the AT-rich, 1.1 megabase (Mb) Mycoplasma mycoides genome in yeast encouraged us to explore similar methods with Prochlorococcus. Prochlorococcus MED4 has an AT-rich genome, with a GC content of 30.8%, similar to that of Saccharomyces cerevisiae (38%), and contains abundant yeast replication origin consensus sites (ACS) evenly distributed around its 1.66 Mb genome. Unlike Mycoplasma cells, which use the UGA codon for tryptophane, Prochlorococcus uses the standard genetic code. Despite this, we observed no toxic effects of several partial and 15 whole Prochlorococcus MED4 genome clones in S. cerevisiae. Sequencing of a Prochlorococcus genome purified from yeast identified 14 single base pair missense mutations, one frameshift, one single base substitution to a stop codon and one dinucleotide transversion compared to the donor genomic DNA. We thus provide evidence of transformation, replication and maintenance of this 1.66 Mb intact bacterial genome in S. cerevisiae.
Source: Tagwerker, C., Dupont, C. L., Karas, B. J., Ma, L., Chuang, R.-Y., Benders, G. A., … Hutchison, C. A. (2012). Sequence analysis of a complete 1.66 Mb Prochlorococcus marinus MED4 genome cloned in yeast. Nucleic Acids Research, 40(20), 10375–10383. doi:10.1093/nar/gks823
Source: Tagwerker, C., Dupont, C. L., Karas, B. J., Ma, L., Chuang, R.-Y., Benders, G. A., … Hutchison, C. A. (2012). Sequence analysis of a complete 1.66 Mb Prochlorococcus marinus MED4 genome cloned in yeast. Nucleic Acids Research, 40(20), 10375–10383. doi:10.1093/nar/gks823
Progress in studying Prochlorococcus from these different perspectives has been hampered by our inability to develop a robust genetic system. Although we and others have worked on this over many years, very limited progress has been made. Possible reasons behind these failures include the cell’s slow growth rate (roughly one doubling per day under optimal laboratory conditions), their reluctance to grow axenically on solid media, specific media requirements [47, 48], sensitivity to trace metal contamination [49, 50] and reactive oxygen species [51], unusual membrane composition [52], and their apparent reduced homologous recombination potential [53]. From our perspective, a significant impediment to rapid progress is that investigators generally do not publish negative results or partial progress; as a result, considerable time is wasted by others in rediscovering what does not work. Thus, before describing our progress, we first report experiments that failed. We initially tried to build upon the only reported successful (albeit inefficient) Prochlorococcus genetic transformation method, based on conjugation from an E. coli donor strain [54]. This method proved unsuccessful in our hands, probably because the six month multi-step procedure is vulnerable to user variability. From there we moved on to explore the use of electroporation and were able to find appropriate conditions for delivering DNA into living Prochlorococcus cells – developing a method that should be applicable to a wide range of microorganisms. The next step will be to develop genetic tools compatible with the host that will also bypass the cell’s defences against exogenous DNA, which are abundant in cyanobacteria [55].
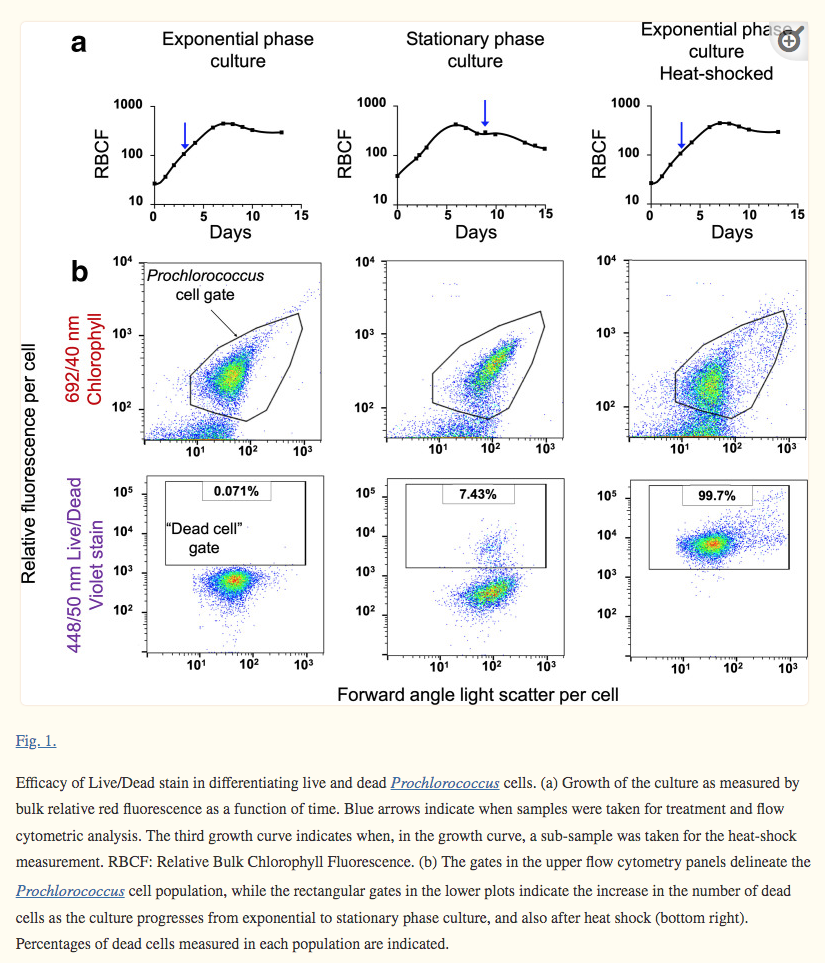
We report the results of some initial attempts, in which transposomes were delivered into Prochlorococcus cells. To avoid the pursuit of unproductive paths, we report here what has not worked in our hands, as well as our progress developing a method to screen the most efficient electroporation parameters for optimal DNA delivery into Prochlorococcus cells. We also report a novel protocol for obtaining axenic colonies and a new method for differentiating live and dead cells (Fig. 1):
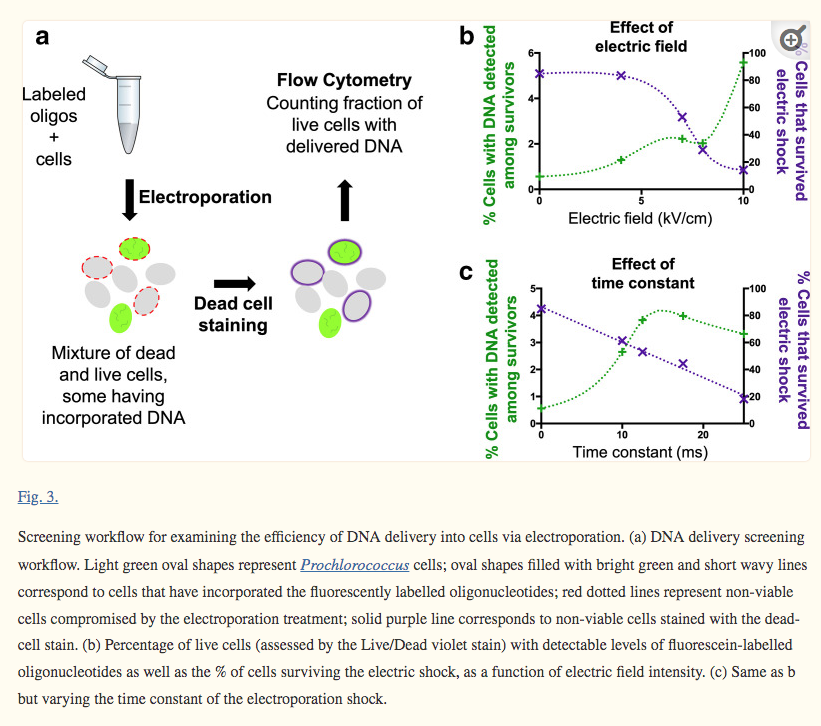
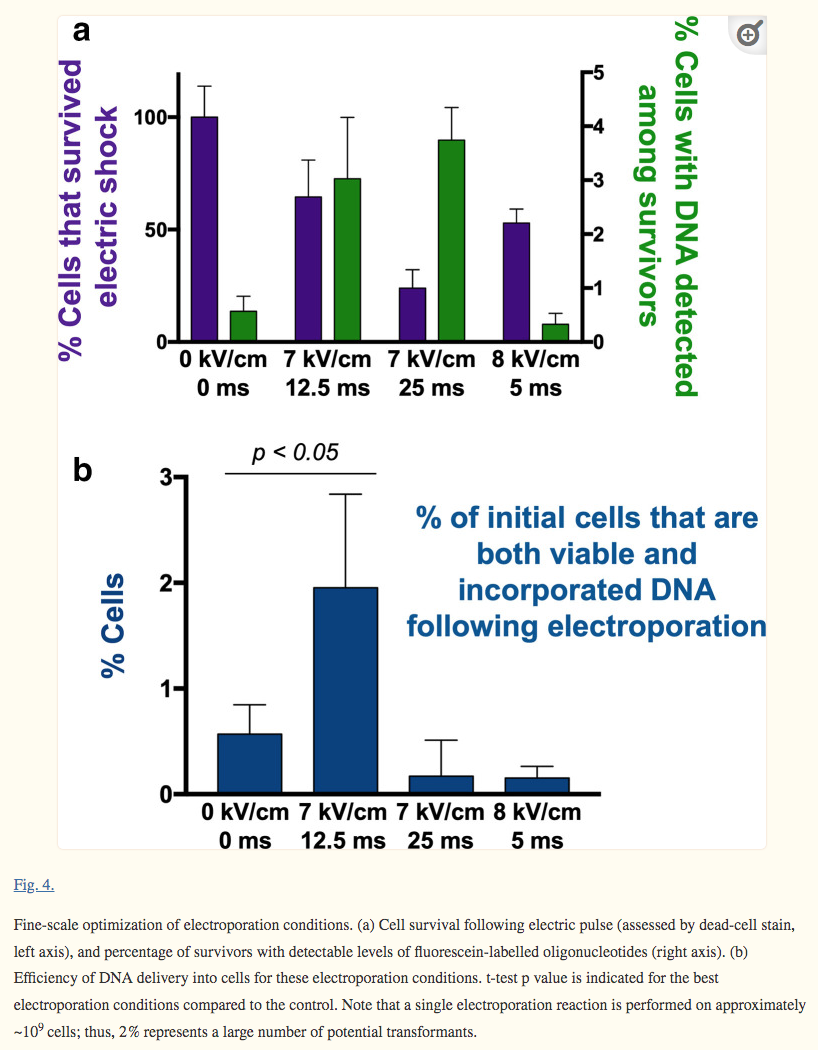
The electroporation method can be used to optimize DNA delivery into any bacterium, making it a useful tool for advancing transformation systems in other genetically recalcitrant microorganisms (Fig. 3 and 4):
Source: Laurenceau, R., Bliem, C., Osburne, M. S., Becker, J. W., Biller, S. J., Cubillos-Ruiz, A., & Chisholm, S. W. (2020). Toward a genetic system in the marine cyanobacterium Prochlorococcus. Access Microbiology, 2(4). https://doi.org/10.1099/ACMI.0.000107
Source: Laurenceau, R., Bliem, C., Osburne, M. S., Becker, J. W., Biller, S. J., Cubillos-Ruiz, A., & Chisholm, S. W. (2020). Toward a genetic system in the marine cyanobacterium Prochlorococcus. Access Microbiology, 2(4). https://doi.org/10.1099/ACMI.0.000107
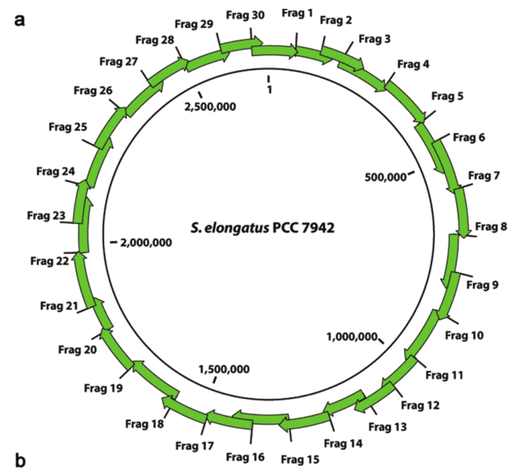
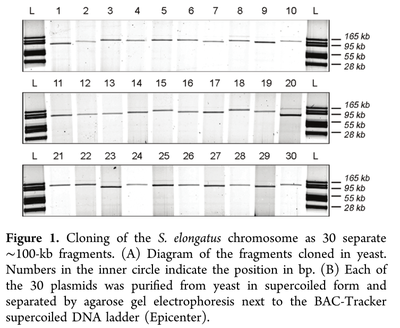
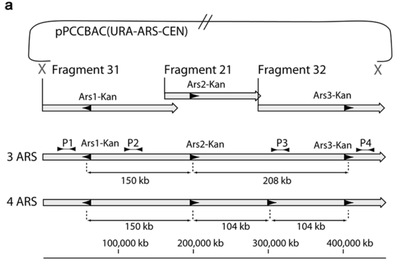
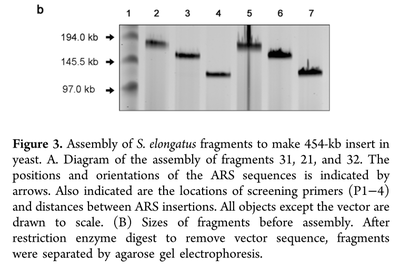
The genome of Synechococcus elongatus PCC 7942, which has an average G+C content of 55%, has been cloned in multiple, separate segments into yeast. None of the regions of the chromosome (approximately 100 kilobase - kb - fragments) are toxic to yeast, however S. elongatus DNA fragments over ∼200 kb are not stably maintained. DNA constructs with a total size under 200 kb could be readily assembled, even with 62 kb of overlapping sequence between pieces. Addition of yeast origins of replication throughout allowed to increase the total size of DNA that could be assembled in yeast by homologous recombination to at least 454 kb. Cloning strategies utilizing yeast recombination with large, high G+C prokaryotic sequences should include yeast origins of replication as a part of the design process.
Source: Noskov, V. N., Karas, B. J., Young, L., Chuang, R.-Y., Gibson, D. G., Lin, Y.-C., … Weyman, P. D. (2012). Assembly of Large, High G+C Bacterial DNA Fragments in Yeast. ACS Synthetic Biology, 120521135004008. doi:10.1021/sb3000194
Source: Noskov, V. N., Karas, B. J., Young, L., Chuang, R.-Y., Gibson, D. G., Lin, Y.-C., … Weyman, P. D. (2012). Assembly of Large, High G+C Bacterial DNA Fragments in Yeast. ACS Synthetic Biology, 120521135004008. doi:10.1021/sb3000194
Genetic modifications in eukaryotic algae:
In a 2011 publication O'neill et al. from Sapphire Energy demonstrate a system for cloning and modifying the chloroplast genome from the green alga, Chlamydomonas reinhardtii. Through extensive use of sequence stabilization strategies, the ex vivo genome is assembled in yeast from a collection of overlapping fragments. The assembled genome is then moved into bacteria for large-scale preparations and transformed into C. reinhardtii cells:
Transformation protocol (typical protocol for transformation of gene and genome fragments into algae): Cells were grown in either TAP or HSM medium and in liquid or on agar plates (23,24). Strains were grown at 28°C, in a 5% CO2 chamber, with 150 uE of light. For transformation, algae cells were grown to 10e6 cells/ml in TAP, collected by centrifugation, and resuspended to 10e8 cells/ml in TAP. A 500 ml aliquot of concentrated culture was spread on either TAP agar plates supplemented with 100 ug/ml of Kanamycin or on HSM agar plates. DNA was bound to 1 micrometer gold particles (Seashell Technology; La Jolla, CA, USA) according to the manufacturers protocol and prepared and bombarded by a Helios Gene Gun (Bio-Rad; Hercules, CA, USA) according to the manufacturer’s instructions. Gold particles were shot using 300 psi of Helium at a distance of 6 cm from the plate.
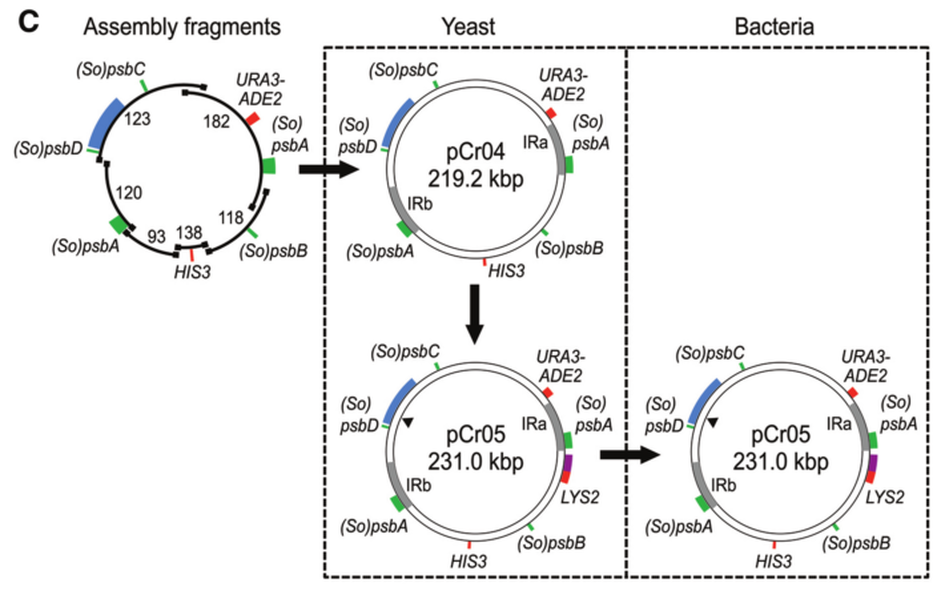
Flow diagram of genome assembly and maintenance in yeast and bacteria. Each arrow represents a transformation step to assemble or modify the genome in yeast or to transfer the genome from yeast to bacteria. In all cases, green boxes indicate PSII-encoding genes, red boxes indicate yeast selection markers, blue boxes indicate the hybrid vector elements from pTrp-34, purple boxes indicate bacterial F-factor replication elements, and gray boxes indicate the large, inverted repeats. Numbers adjacent to each assembly fragment correspond to the pSC vector from which they were liberated. The black triangle in pCr05 indicates the unique AsiSI restriction site.
Source: O’Neill, B. M., Mikkelson, K. L., Gutierrez, N. M., Cunningham, J. L., Wolff, K. L., Szyjka, S. J., … Mendez, M. J. (2011). An exogenous chloroplast genome for complex sequence manipulation in algae. Nucleic Acids Research, 40(6), 2782–2792. doi:10.1093/nar/gkr1008
Source: O’Neill, B. M., Mikkelson, K. L., Gutierrez, N. M., Cunningham, J. L., Wolff, K. L., Szyjka, S. J., … Mendez, M. J. (2011). An exogenous chloroplast genome for complex sequence manipulation in algae. Nucleic Acids Research, 40(6), 2782–2792. doi:10.1093/nar/gkr1008
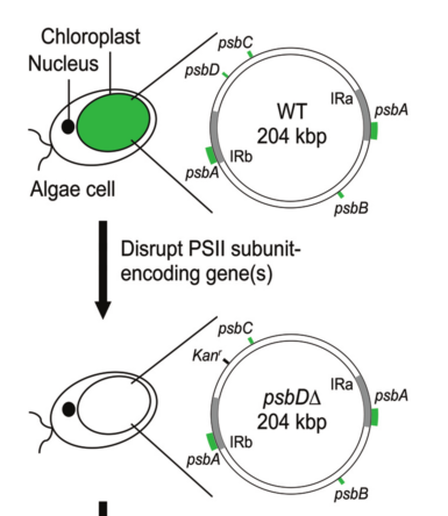
To characterize in vivo function of the cloned C. reinhardtii chloroplast genome, a strategy was used that relies on the restoration of photosynthesis as the primary selection (Figure 2). O'neill et al. first created a collection of photosynthetically deficient algae strains into which the genome could be transformed (see Supplementary Figure S4 and ‘Materials and Methods’ section in publication). Each strain in the collection contains a knockout in the coding region of psbA, psbB, psbC or psbD; these modifications render them unable to grow without the addition of organic carbon to compensate for their inability to perform photosynthesis (Supplementary Figure S4). Each strain in the collection is also complemented by transformation with its corresponding wild-type locus (Supplementary Figure S5), demonstrating that they are all transformable.
The researchers went on to use this system to substitute genes encoding core subunits of the photosynthetic apparatus with orthologs from a related alga, Scenedesmus obliquus. Once transformed into algae, the substituted genome recombines with the endogenous genome, resulting in a hybrid plastome comprising modifications in disparate loci. The in vivo function of the genomes described herein demonstrates that simultaneous engineering of multiple sites within the chloroplast genome is now possible.
The researchers went on to use this system to substitute genes encoding core subunits of the photosynthetic apparatus with orthologs from a related alga, Scenedesmus obliquus. Once transformed into algae, the substituted genome recombines with the endogenous genome, resulting in a hybrid plastome comprising modifications in disparate loci. The in vivo function of the genomes described herein demonstrates that simultaneous engineering of multiple sites within the chloroplast genome is now possible.
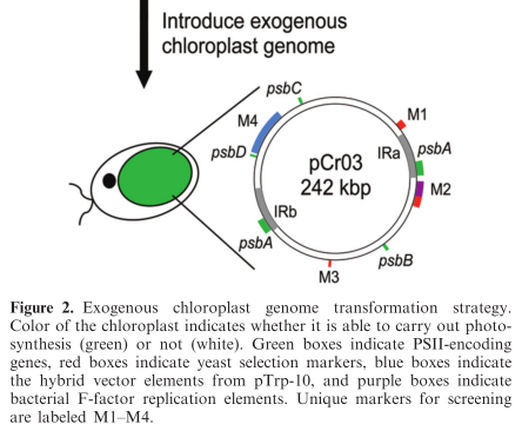
Taken together, all of the in vivo data demonstrate that the cloned C. reinhardtii chloroplast genome provides a platform for simultaneous multi-site modification of the photosynthetic apparatus. Replaced sequences can be native or orthologous genes, traditionally cloned or synthetically created. These results also show that this approach can be used to simultaneously modify at least six disparate regions in the chloroplast genome in vivo.
Source: O’Neill, B. M., Mikkelson, K. L., Gutierrez, N. M., Cunningham, J. L., Wolff, K. L., Szyjka, S. J., … Mendez, M. J. (2011). An exogenous chloroplast genome for complex sequence manipulation in algae. Nucleic Acids Research, 40(6), 2782–2792. doi:10.1093/nar/gkr1008
Source: O’Neill, B. M., Mikkelson, K. L., Gutierrez, N. M., Cunningham, J. L., Wolff, K. L., Szyjka, S. J., … Mendez, M. J. (2011). An exogenous chloroplast genome for complex sequence manipulation in algae. Nucleic Acids Research, 40(6), 2782–2792. doi:10.1093/nar/gkr1008
Cloning of 2 chromosomes extracted from the model diatom Phaeodactylum tricornutum:
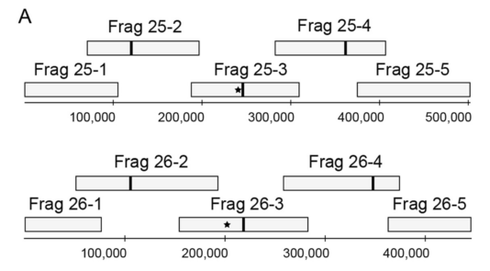
It has been previously demonstrated that cloning in yeast of large (> ~150 kb), high G+C (55%) prokaryotic DNA fragments was improved by addition of yeast replication origins every ~100 kb (see Noskov, V. N., Karas, B. J., Young, L., Chuang, R.-Y., Gibson, D. G., Lin, Y.-C., … Weyman, P. D. (2012). Assembly of Large, High G+C Bacterial DNA Fragments in Yeast. ACS Synthetic Biology). Conversely, low GC DNA is stable (up to at least 1.8 Mb) without adding supplemental yeast origins. It has not been previously tested whether addition of yeast replication origins similarly improves the yeast-based cloning of large (>150 kb) eukaryotic DNA with moderate GC content. The model diatom Phaeodactylum tricornutum has an average GC content of 48% and a 27.4 Mb genome sequence that has been assembled into chromosome-sized scaffolds making it an ideal test case for assembly and maintenance of eukaryotic chromosomes in yeast.
A modified chromosome assembly technique in which eukaryotic chromosomes as large as ~500 kb can be assembled from cloned ~100kb fragments has been developed. This technique was used to clone fragments spanning P. tricornutum chromosomes 25 and 26 and to assemble these fragments into single, chromosome-sized molecules. The researchers found that addition of yeast replication origins improved the cloning, assembly, and maintenance of the large chromosomes in yeast. Furthermore, purification of the fragments to be assembled by electroelution greatly increased assembly efficiency.
To summarize, entire eukaryotic chromosomes can be successfully cloned, maintained, and manipulated in yeast. These results highlight the improvement in assembly and maintenance afforded by including yeast replication origins in eukaryotic DNA with moderate GC content (48%). They also highlight the increased efficiency of assembly that can be achieved by purifying fragments before assembly.
A modified chromosome assembly technique in which eukaryotic chromosomes as large as ~500 kb can be assembled from cloned ~100kb fragments has been developed. This technique was used to clone fragments spanning P. tricornutum chromosomes 25 and 26 and to assemble these fragments into single, chromosome-sized molecules. The researchers found that addition of yeast replication origins improved the cloning, assembly, and maintenance of the large chromosomes in yeast. Furthermore, purification of the fragments to be assembled by electroelution greatly increased assembly efficiency.
To summarize, entire eukaryotic chromosomes can be successfully cloned, maintained, and manipulated in yeast. These results highlight the improvement in assembly and maintenance afforded by including yeast replication origins in eukaryotic DNA with moderate GC content (48%). They also highlight the increased efficiency of assembly that can be achieved by purifying fragments before assembly.
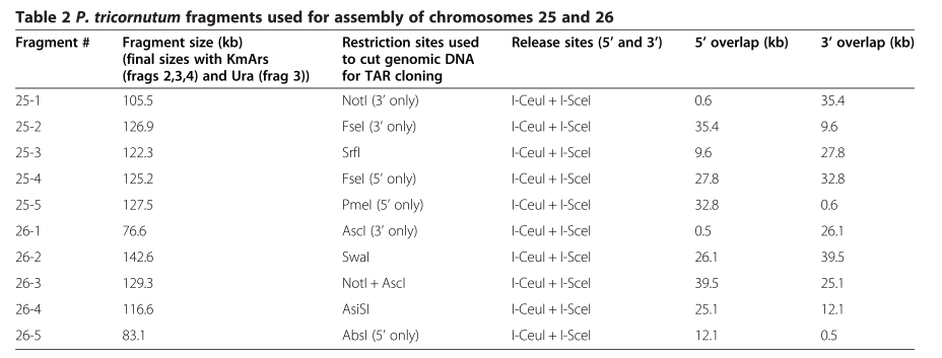

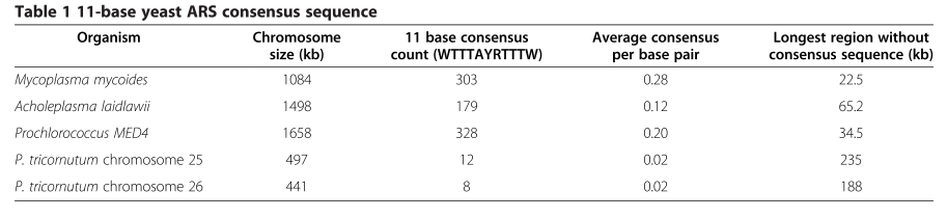
DNA with low GC content will have a greater chance of having AT-rich sites that function as yeast replication origins while DNA of higher average GC content will have statistically fewer of these sites. Insertion of known replication origins every 100-150 kb facilitates cloning of large DNA fragments into S. cerevisiae.
Source: Karas, B. J., Molparia, B., Jablanovic, J., Hermann, W. J., Lin, Y.-C., Dupont, C. L., … Weyman, P. D. (2013). Assembly of eukaryotic algal chromosomes in yeast. Journal of Biological Engineering, 7(1), 30. doi:10.1186/1754-1611-7-30