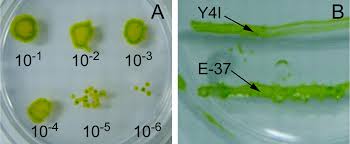
Individual species:
Eukaryotic Algae:
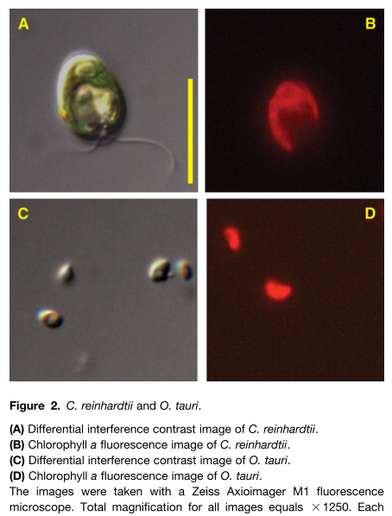
Measuring just under 1 micrometer in diameter, Ostreococcus (left Figure, bottom panel) claims the title of smallest known free-living eukaryote (Courties et al., 1994). The nuclear genomes of O. tauri (Derelle et al., 2006) and O. lucimarinus (Palenik et al., 2007) are also small, weighing in at only 12.6 and 13.2 megabases (Mb), respectively, and they are predicted to have 7892 and 7651 nuclear genes packed into 20 and 21 chromosomes, respectively. These genomes contain considerably more genes than the relatively simple oxygenic prokaryote Prochlorococcus (1.7 Mb and approximately 1900 protein-coding genes; Dufresne et al., 2003), but they contain only slightly more genes than are found in more complex cyanobacteria like Trichodesmium (7.75 Mb and approximately 6300 total genes). The structure of Ostreococcus genes illustrates the outcome of severe genome compaction. Only one-quarter of gene models in O. tauri have more than one predicted intron, and for the genome as a whole, it is predicted that there are only 1.57 exons per gene, with an average transcript size of 1257 bp. O. lucimarinus has only 20% of predicted genes containing multiple introns with similar exon number (1.27 per gene) and similar transcript size (1234 bp) as its kin.
Source: Peers, G., & Niyogi, K. K. (2008). Pond scum genomics: the genomes of Chlamydomonas and Ostreococcus. The Plant Cell, 20(3), 502–7. doi:10.1105/tpc.107.056556
Source: Peers, G., & Niyogi, K. K. (2008). Pond scum genomics: the genomes of Chlamydomonas and Ostreococcus. The Plant Cell, 20(3), 502–7. doi:10.1105/tpc.107.056556
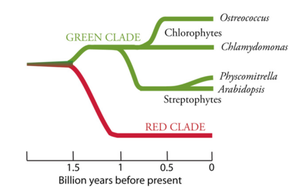
By contrast, the 17 linkage groups of Chlamydomonas (top panel, Figure above) are seemingly awash in excess genetic code. The draft sequence (Merchant et al., 2007) has 15,143 predicted genes over 121 Mb, and the genes appear riddled with introns (average of 7.3 per gene) that extend the average transcript length of 1580 bp to an average gene length of 4312 bp (see Table 1 in Merchant et al. [2007] for a detailed comparison). Beyond the contrasting patterns of genome structure, Ostreococcus and Chlamydomonas exhibit unusually high GC contents. The GC content of the O. tauri genome is 58%, while the GC content of Chlamydomonas is a whopping 64% (68% in coding regions!). Another notable feature of the Chlamydomonas genome is its high density of repetitive DNA sequences. Many small silencing RNAs with sequence similarity to transposable elements (TEs) have been found recently in the Chlamydomonas genome (Molnar et al., 2007), along with the first microRNAs described for a unicellular organism (Molnar et al., 2007; Zhao et al., 2007). The green clade of photosynthetic eukaryotes includes Chlorophytes (green algae) and Streptophytes (plants and charophycean green algae). Ostreococcus is a member of the class Prasinophyceae, whereas Chlamydomonas is classified in the Chlorophyceae. The green and red clades share a common ancestor: a heterotrophic eukaryote that en- gulfed a cyanobacterium to generate a plastid through primary endosymbiosis. The scale represents approximate times of divergence as taken from Yoon et al. (2004). The time of divergence of prasinophytes from other Chlorophytes is not well characterized.
Source: Peers, G., & Niyogi, K. K. (2008). Pond scum genomics: the genomes of Chlamydomonas and Ostreococcus. The Plant Cell, 20(3), 502–7. doi:10.1105/tpc.107.056556
Source: Peers, G., & Niyogi, K. K. (2008). Pond scum genomics: the genomes of Chlamydomonas and Ostreococcus. The Plant Cell, 20(3), 502–7. doi:10.1105/tpc.107.056556
The marine pennate diatom Phaeodactylum is often studied in laboratories because of the superior genetic resources available for this diatom (eg, genetic transformation, 100,000 ESTs), a whole genome sequence has been generated and because it has been used in laboratory-based studies of diatom physiology for several decades. Although not considered to be of great ecological significance, it has been found in several locations around the world, typically in coastal areas with wide fluctuations in salinity. Unlike other diatoms it can exist in different morphotypes, and changes in cell shape can be stimulated by environmental conditions. This feature can be used to explore the molecular basis of cell shape control and morphogenesis. Furthermore the species can grow in the absence of silicon, and the biogenesis of silicified frustules is facultative, thereby providing opportunities for experimental exploration of silicon-based nanofabrication in diatoms.
The marine pennate diatom Phaeodactylum is often studied in laboratories because of the superior genetic resources available for this diatom (eg, genetic transformation, 100,000 ESTs), a whole genome sequence has been generated and because it has been used in laboratory-based studies of diatom physiology for several decades. Although not considered to be of great ecological significance, it has been found in several locations around the world, typically in coastal areas with wide fluctuations in salinity. Unlike other diatoms it can exist in different morphotypes, and changes in cell shape can be stimulated by environmental conditions. This feature can be used to explore the molecular basis of cell shape control and morphogenesis. Furthermore the species can grow in the absence of silicon, and the biogenesis of silicified frustules is facultative, thereby providing opportunities for experimental exploration of silicon-based nanofabrication in diatoms.
As a diatom Phaeodactylum is characterized by a photosynthetic apparatus that contains xanthophylls (fucoxanthin) and chlorophylls a/c as dominant light-harvesting pigments (Grossman et al., 1995). Diatoms and other chromophytic algae also possess ‘complex plastids’ delineated by four distinct membranes (Gibbs, 1979). This feature has been proposed to reflect the origin of chromophytic plastids by a secondary endocytobiotic event: that is, engulfment of a eukaryotic alga by a nonphotosynthetic eukaryote that led to permanent residence of the photosynthetic organism within the host cells and subsequent evolution of a plastid. The ancestral chromophytic plastid is thought to be related to red algae (Martin et al., 1998), which themselves arose by primary endocytobiosis (1).
Four membranes delineate the chromophytic plastid. The inner two appear to be homologous to the double membrane envelope of plastids from red algae, green algae and vascular plants. A third unique membrane completely surrounds the plastid envelope and is thought to represent the former plasma membrane of the endosymbiont, while the outermost membrane may have evolved from the vacuolar/plasma membrane of the host organism. This outermost membrane has been observed to be continuous with the endoplasmic reticulum (Gibbs, 1981; Ishida et al., 2000). The portion of this ER-like membrane directly adjacent to plastids (and which appears to completely encase the plastid) is commonly referred to as the chloroplast ER or CER (Bouck, 1965).
Four membranes delineate the chromophytic plastid. The inner two appear to be homologous to the double membrane envelope of plastids from red algae, green algae and vascular plants. A third unique membrane completely surrounds the plastid envelope and is thought to represent the former plasma membrane of the endosymbiont, while the outermost membrane may have evolved from the vacuolar/plasma membrane of the host organism. This outermost membrane has been observed to be continuous with the endoplasmic reticulum (Gibbs, 1981; Ishida et al., 2000). The portion of this ER-like membrane directly adjacent to plastids (and which appears to completely encase the plastid) is commonly referred to as the chloroplast ER or CER (Bouck, 1965).
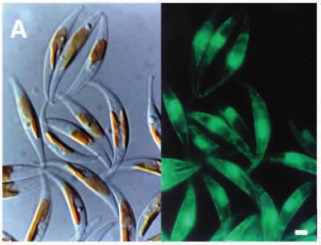
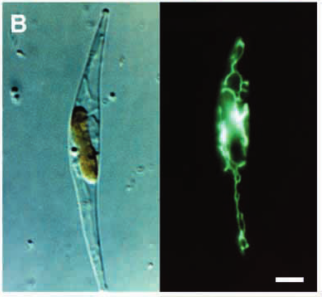
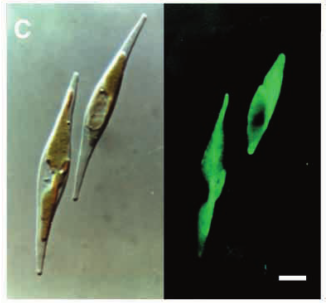
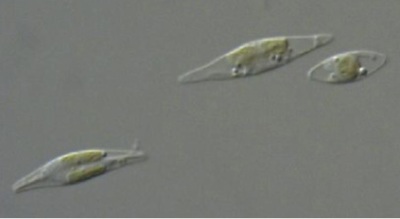
Calcofluor white stained cell walls of Phaeodactylum tricornutum under DIC light microscopy and fluorescence microscopy: the plastid appears red due to excitation of chlorophyll molecules at low fluorescence excitation wavelengths and brown in the light microscope due to xanthophylls (fucoxanthin) characteristic of the Phaeophyceae (brown algae):
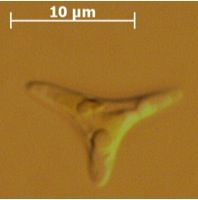
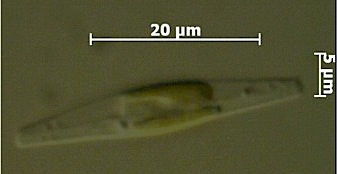
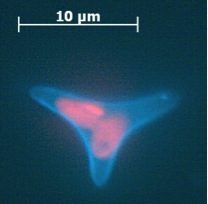
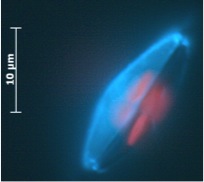
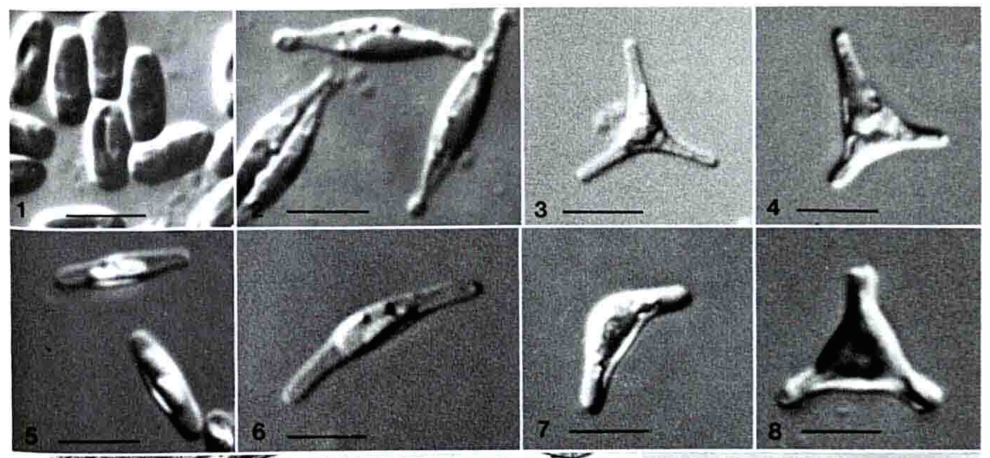
The ultrastructure of the oval, fusiform and triradiate morphotypes of Phaeodactylum tricornutum Bohlin. The organization and structure of the cytoplasmic organelles is similar in all three morphotypes, except that the vacuoles occupy the extra volume created by the arms of the fusiform and triradiate cells. The frustule in fusiform and triradiate cells is organic; in the oval type it may be organic or one of the valves may have a silica frustule surrounded by an organic wall. In all cells, the organic cell wall has up to 10 silica bands (13 nm wide) embedded in its surface in the girdle region, lacks girdle bands, and has an outer corrugated cell wall layer, except in the girdle region. Cell division, organic wall formation and silica deposition have been researched in detail. Four types of oval cells are also described (2).
Scale bars = 10 micrometers
1.) Oval or ovoid morphotype 2.) Fusiform morphotype 3.) "long-armed" triradiate 4.) Triradiate with unequal arms 5.) "Long" oval/ovoid 6.) "Bent" (lunate) fusiform 7.) "Boomerang" fusiform 8.) "short-armed" triradiate.
Sources:
Scale bars = 10 micrometers
1.) Oval or ovoid morphotype 2.) Fusiform morphotype 3.) "long-armed" triradiate 4.) Triradiate with unequal arms 5.) "Long" oval/ovoid 6.) "Bent" (lunate) fusiform 7.) "Boomerang" fusiform 8.) "short-armed" triradiate.
Sources:
- Apt, K. E., Zaslavkaia, L., Lippmeier, J. C., Lang, M., Kilian, O., Wetherbee, R., … Kroth, P. G. (2002). In vivo characterization of diatom multipartite plastid targeting signals. Journal of Cell Science, 115(21), 4061.
- Borowitzka, M. A., & Volcani, B. E. (1978). THE POLYMORPHIC DIATOM PHAEODACTYLUM TRICORNUTUM: ULTRASTRUCTURE OF ITS MORPHOTYPES 2. Journal of Phycology, 14(1), 10–21.
Prokaryotic Algae:
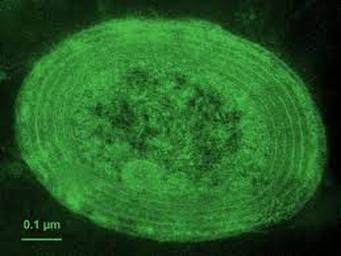
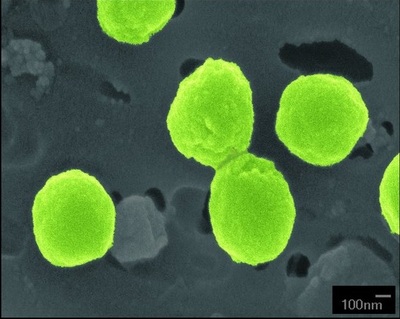
Prochlorococcus is a marine cyanobacterium that numerically dominates the mid-latitude oceans, and is the smallest known oxygenic phototroph. All isolates described thus far can be assigned to either a tightly clustered high-light (HL) adapted clade, or a more divergent low-light (LL) adapted group. They are closely related to, but distinct from, marine Synechococcus. The genomes of 12 strains have been sequenced and they range in size from 1.6 to 2.6 Mbp. They represent diverse lineages, spanning the rRNA diversity (97 to 99.93% similarity) of cultured representatives of this group. Our analyses of these genomes inform our understanding of how adaptation occurs in the oceans along gradients of light, nutrients, and other environmental factors, providing essential context for interpreting rapidly expanding metagenomic datasets. Click for Prochlorococcus genomes list. Click on images of Prochlorococcus microscopy pictures or saltwater medium agar plate and liquid cultures below for more information.
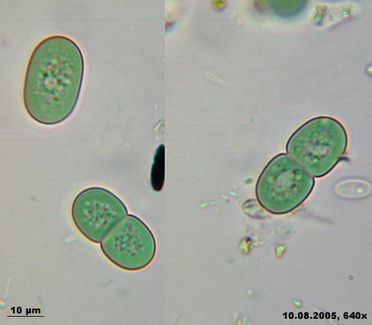
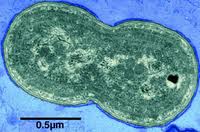
The freshwater cyanobacterium Synechococcus elongatus PCC 7942 holds a special place in the development of cyanobacterial molecular genetics. It was this isolate, previously known as Anacystis nidulans R2, that was the first cyanobacterium demonstrated to be reliably transformable by exogenously added DNA (32). Over the decades, many genetic tools have been developed for S. elongatus, and applied to other transformable cyanobacteria (1, 9, 16). Strain PCC 7942 is very closely related to, and considered the same species as, PCC 6301, which is the type strain for S. elongatus (13) and the first accession in the Pasteur Culture Collection (http://www.pasteur.fr/recherche/banques/PCC/). It is an obligate photoautotroph (13), with a genome of approximately 2.7 Mb (15). Two plasmids (approximately 8 and 46 kb) have been sequenced entirely (GenBank accessions S89470 and AF441790, respectively). S. elongatus has been extensively studied by a multi-national research community with respect to: acquisition of inorganic carbon (2, 8, 20, 22, 33), transport and regulation of nitrogen compounds (19, 23, 36), response to iron deprivation (3, 7, 21, 24, 37), acclimation to a variety of nutrient stresses (5, 6, 12, 27, 28, 31, 35), and adaptation to environmental variations in temperature (11, 14, 25, 26, 30) and light intensity (4, 17, 18, 29, 31, 34, 35). In recent years, S. elongatus PCC 7942 has again taken a pioneering position, as the only developed model system for exploring the mechanism of a prokaryotic circadian clock (10). A companion project to the JGI draft sequence is a functional genomics project (http://www.bio.tamu.edu/synecho/index.html) that aims to inactivate each gene in the genome and identify all loci that are important for circadian rhythms of gene expression in the organism.
Source: JGI - DOE Joint Genome Institute
1. Andersson CA, Tsinoremas NF, Shelton J, Lebedeva NV, Yarrow J, et al. 2000. Application of bioluminescence to the study of circadian rhythms in cyanobacteria. Methods Enzymol. 305: 527-42
2. Bonfil DJ, Ronen-Tarazi M, Sultemeyer D, Lieman-Hurwitz J, Schatz D, Kaplan A. 1998. A putative HCO3- transporter in the cyanobacterium Synechococcus sp. strain PCC 7942. FEBS Lett. 430: 236-40
3. Burnap RL, Troyan T, Sherman LA. 1993. The highly abundant chlorophyll-protein complex of iron-deficient Synechococcus sp. PCC7942 (CP43') is encoded by the isiA gene. Plant Physiol. 103: 893-902
4. Bustos SA, Golden SS. 1992. Light-regulated expression of the psbD gene family in Synechococcus sp. strain PCC 7942: evidence for the role of duplicated psbD genes in cyanobacteria. Mol. Gen. Genet. 232: 221-30
5. Collier JL, Grossman AR. 1992. Chlorosis induced by nutrient deprivation in Synechococcus sp. strain PCC 7942: not all bleaching is the same. J. Bacteriol. 174: 4718-26
6. Collier JL, Grossman AR. 1994. A small polypeptide triggers complete degradation of light-harvesting phycobiliproteins in nutrient-deprived cyanobacteria. Embo J. 13: 1039-47
7. Durham KA, Porta D, McKay RM, Bullerjahn GS. 2003. Expression of the iron-responsive irpA gene from the cyanobacterium Synechococcus sp strain PCC 7942. Arch. Microbiol. 179: 131-4
8. Gabay C, Lieman-Hurwitz J, Hassidim M, Ronen-Tarazi M, Kaplan A. 1998. Modification of topA in Synechococcus sp. PCC 7942 resulted in mutants capable of growing under low but not high concentration of CO2. FEMS Microbiol. Lett. 159: 343-7
9. Golden SS, Brusslan J, Haselkorn R. 1987. Genetic engineering of the cyanobacterial chromosome. Methods Enzymol. 153: 215-31
10. Golden SS, Johnson CH, Kondo T. 1998. The cyanobacterial circadian system: a clock apart. Curr. Opin. Microbiol. 1: 669-73
11. Gombos Z, Kanervo E, Tsvetkova N, Sakamoto T, Aro EM, Murata N. 1997. Genetic enhancement of the ability to tolerate photoinhibition by introduction of unsaturated bonds into membrane glycerolipids. Plant Physiol. 115: 551-9
12. Green LS, Laudenbach DE, Grossman AR. 1989. A region of a cyanobacterial genome required for sulfate transport. Proc. Natl. Acad. Sci. U S A 86: 1949-53
13. Herdman M, Castenholz RW, Iteman I, Waterbury JB, Rippka R. 2001. Subsection I: (Formerly Chroococcales Wettstein 1924, emend. Rippka, Deruelles, Waterbury, Herdman and Stanier 1979). In Bergey's Manual of Systematic Bacteriology, ed. DR Boone, RW Castenholz, GM Garrity, pp. 721. New York: Springler-Verlag
14. Hossain MM, Nakamoto H. 2002. HtpG plays a role in cold acclimation in cyanobacteria. Curr. Microbiol. 44: 291-6
15. Kaneko T, Matsubayashi T, Sugita M, Sugiura M. 1996. Physical and gene maps of the unicellular cyanobacterium Synechococcus sp. strain PCC6301 genome. Plant Mol. Biol. 31: 193-201
16. Kuhlemeier CJ, van Arkel GA. 1987. Host-vector systems for gene cloning in cyanobacteria. Methods Enzymol. 153: 199-215
17. Kulkarni RD, Golden SS. 1997. mRNA stability is regulated by a coding-region element and the unique 5' untranslated leader sequences of the three Synechococcus psbA transcripts. Mol Microbiol 24: 1131-42
18. Kulkarni RD, Schaefer MR, Golden SS. 1992. Transcriptional and posttranscriptional components of psbA response to high light intensity in Synechococcus sp. strain PCC 7942. J Bacteriol 174: 3775-81
19. Lee HM, Vazquez-Bermudez MF, de Marsac NT. 1999. The global nitrogen regulator NtcA regulates transcription of the signal transducer PII (GlnB) and influences its phosphorylation level in response to nitrogen and carbon supplies in the cyanobacterium Synechococcus sp. strain PCC 7942. J. Bacteriol. 181: 2697-702
20. Maeda S, Price GD, Badger MR, Enomoto C, Omata T. 2000. Bicarbonate binding activity of the CmpA protein of the cyanobacterium Synechococcus sp. strain PCC 7942 involved in active transport of bicarbonate. J. Biol. Chem. 275: 20551-5
21. Michel KP, Pistorius EK, Golden SS. 2001. Unusual regulatory elements for iron deficiency induction of the idiA gene of Synechococcus elongatus PCC 7942. J. Bacteriol. 183: 5015-24
22. Omata T, Gohta S, Takahashi Y, Harano Y, Maeda S. 2001. Involvement of a CbbR homolog in low CO2-induced activation of the bicarbonate transporter operon in cyanobacteria. J. Bacteriol. 183: 1891-8
23. Sakamoto T, Bryant DA. 1999. Nitrate transport and not photoinhibition limits growth of the freshwater cyanobacterium Synechococcus species PCC 6301 at low temperature. Plant Physiol. 119: 785-94
24. Sandstrom S, Ivanov AG, Park YI, Oquist G, Gustafsson P. 2002. Iron stress responses in the cyanobacterium Synechococcus sp. PCC7942. Physiol. Plant 116: 255-63
25. Sane PV, Ivanov AG, Sveshnikov D, Huner NP, Oquist G. 2002. A transient exchange of the photosystem II reaction center protein D1:1 with D1:2 during low temperature stress of Synechococcus sp. PCC 7942 in the light lowers the redox potential of QB. J. Biol. Chem. 277: 32739-45
26. Sarcina M, Tobin MJ, Mullineaux CW. 2001. Diffusion of phycobilisomes on the thylakoid membranes of the cyanobacterium Synechococcus 7942. Effects of phycobilisome size, temperature, and membrane lipid composition. J. Biol. Chem. 276: 46830-4
27. Sauer J, Gorl M, Forchhammer K. 1999. Nitrogen starvation in Synechococcus PCC 7942: involvement of glutamine synthetase and NtcA in phycobiliprotein degradation and survival. Arch. Microbiol. 172: 247-55
28. Sauer J, Schreiber U, Schmid R, Volker U, Forchhammer K. 2001. Nitrogen starvation-induced chlorosis in Synechococcus PCC 7942. Low-level photosynthesis as a mechanism of long-term survival. Plant Physiol. 126: 233-43
29. Schaefer MR, Golden SS. 1989. Light availability influences the ratio of two forms of D1 in cyanobacterial thylakoids. J. Biol. Chem. 264: 7412-7
30. Schelin J, Lindmark F, Clarke AK. 2002. The clpP multigene family for the ATP-dependent Clp protease in the cyanobacterium Synechococcus. Microbiology 148: 2255-65
31. Schwarz R, Grossman AR. 1998. A response regulator of cyanobacteria integrates diverse environmental signals and is critical for survival under extreme conditions. Proc. Natl. Acad. Sci. U S A 95: 11008-13
32. Shestakov SV, Khyen NT. 1970. Evidence for genetic transformation in blue-green alga Anacystis nidulans R2. Molec. Gen. Genet. 107: 372-5
33. Tchernov D, Helman Y, Keren N, Luz B, Ohad I, et al. 2001. Passive entry of CO2 and its energy-dependent intracellular conversion to HCO3- in cyanobacteria are driven by a photosystem I-generated deltamuH+. J. Biol. Chem. 276: 23450-5
34. Tsinoremas NF, Schaefer MR, Golden SS. 1994. Blue and red light reversibly control psbA expression in the cyanobacterium Synechococcus sp. strain PCC 7942. J. Biol. Chem. 269: 16143-7
35. van Waasbergen LG, Dolganov N, Grossman AR. 2002. nblS, a gene involved in controlling photosynthesis-related gene expression during high light and nutrient stress in Synechococcus elongatus PCC 7942. J. Bacteriol. 184: 2481-90
36. Vazquez-Bermudez MF, Paz-Yepes J, Herrero A, Flores E. 2002. The NtcA-activated amt1 gene encodes a permease required for uptake of low concentrations of ammonium in the cyanobacterium Synechococcus sp. PCC 7942. Microbiol. 148: 861-9
37. Webb R, Troyan T, Sherman D, Sherman LA. 1994. MapA, an iron-regulated, cytoplasmic membrane protein in the cyanobacterium Synechococcus sp. strain PCC7942. J. Bacteriol. 176: 4906-13
Source: JGI - DOE Joint Genome Institute
1. Andersson CA, Tsinoremas NF, Shelton J, Lebedeva NV, Yarrow J, et al. 2000. Application of bioluminescence to the study of circadian rhythms in cyanobacteria. Methods Enzymol. 305: 527-42
2. Bonfil DJ, Ronen-Tarazi M, Sultemeyer D, Lieman-Hurwitz J, Schatz D, Kaplan A. 1998. A putative HCO3- transporter in the cyanobacterium Synechococcus sp. strain PCC 7942. FEBS Lett. 430: 236-40
3. Burnap RL, Troyan T, Sherman LA. 1993. The highly abundant chlorophyll-protein complex of iron-deficient Synechococcus sp. PCC7942 (CP43') is encoded by the isiA gene. Plant Physiol. 103: 893-902
4. Bustos SA, Golden SS. 1992. Light-regulated expression of the psbD gene family in Synechococcus sp. strain PCC 7942: evidence for the role of duplicated psbD genes in cyanobacteria. Mol. Gen. Genet. 232: 221-30
5. Collier JL, Grossman AR. 1992. Chlorosis induced by nutrient deprivation in Synechococcus sp. strain PCC 7942: not all bleaching is the same. J. Bacteriol. 174: 4718-26
6. Collier JL, Grossman AR. 1994. A small polypeptide triggers complete degradation of light-harvesting phycobiliproteins in nutrient-deprived cyanobacteria. Embo J. 13: 1039-47
7. Durham KA, Porta D, McKay RM, Bullerjahn GS. 2003. Expression of the iron-responsive irpA gene from the cyanobacterium Synechococcus sp strain PCC 7942. Arch. Microbiol. 179: 131-4
8. Gabay C, Lieman-Hurwitz J, Hassidim M, Ronen-Tarazi M, Kaplan A. 1998. Modification of topA in Synechococcus sp. PCC 7942 resulted in mutants capable of growing under low but not high concentration of CO2. FEMS Microbiol. Lett. 159: 343-7
9. Golden SS, Brusslan J, Haselkorn R. 1987. Genetic engineering of the cyanobacterial chromosome. Methods Enzymol. 153: 215-31
10. Golden SS, Johnson CH, Kondo T. 1998. The cyanobacterial circadian system: a clock apart. Curr. Opin. Microbiol. 1: 669-73
11. Gombos Z, Kanervo E, Tsvetkova N, Sakamoto T, Aro EM, Murata N. 1997. Genetic enhancement of the ability to tolerate photoinhibition by introduction of unsaturated bonds into membrane glycerolipids. Plant Physiol. 115: 551-9
12. Green LS, Laudenbach DE, Grossman AR. 1989. A region of a cyanobacterial genome required for sulfate transport. Proc. Natl. Acad. Sci. U S A 86: 1949-53
13. Herdman M, Castenholz RW, Iteman I, Waterbury JB, Rippka R. 2001. Subsection I: (Formerly Chroococcales Wettstein 1924, emend. Rippka, Deruelles, Waterbury, Herdman and Stanier 1979). In Bergey's Manual of Systematic Bacteriology, ed. DR Boone, RW Castenholz, GM Garrity, pp. 721. New York: Springler-Verlag
14. Hossain MM, Nakamoto H. 2002. HtpG plays a role in cold acclimation in cyanobacteria. Curr. Microbiol. 44: 291-6
15. Kaneko T, Matsubayashi T, Sugita M, Sugiura M. 1996. Physical and gene maps of the unicellular cyanobacterium Synechococcus sp. strain PCC6301 genome. Plant Mol. Biol. 31: 193-201
16. Kuhlemeier CJ, van Arkel GA. 1987. Host-vector systems for gene cloning in cyanobacteria. Methods Enzymol. 153: 199-215
17. Kulkarni RD, Golden SS. 1997. mRNA stability is regulated by a coding-region element and the unique 5' untranslated leader sequences of the three Synechococcus psbA transcripts. Mol Microbiol 24: 1131-42
18. Kulkarni RD, Schaefer MR, Golden SS. 1992. Transcriptional and posttranscriptional components of psbA response to high light intensity in Synechococcus sp. strain PCC 7942. J Bacteriol 174: 3775-81
19. Lee HM, Vazquez-Bermudez MF, de Marsac NT. 1999. The global nitrogen regulator NtcA regulates transcription of the signal transducer PII (GlnB) and influences its phosphorylation level in response to nitrogen and carbon supplies in the cyanobacterium Synechococcus sp. strain PCC 7942. J. Bacteriol. 181: 2697-702
20. Maeda S, Price GD, Badger MR, Enomoto C, Omata T. 2000. Bicarbonate binding activity of the CmpA protein of the cyanobacterium Synechococcus sp. strain PCC 7942 involved in active transport of bicarbonate. J. Biol. Chem. 275: 20551-5
21. Michel KP, Pistorius EK, Golden SS. 2001. Unusual regulatory elements for iron deficiency induction of the idiA gene of Synechococcus elongatus PCC 7942. J. Bacteriol. 183: 5015-24
22. Omata T, Gohta S, Takahashi Y, Harano Y, Maeda S. 2001. Involvement of a CbbR homolog in low CO2-induced activation of the bicarbonate transporter operon in cyanobacteria. J. Bacteriol. 183: 1891-8
23. Sakamoto T, Bryant DA. 1999. Nitrate transport and not photoinhibition limits growth of the freshwater cyanobacterium Synechococcus species PCC 6301 at low temperature. Plant Physiol. 119: 785-94
24. Sandstrom S, Ivanov AG, Park YI, Oquist G, Gustafsson P. 2002. Iron stress responses in the cyanobacterium Synechococcus sp. PCC7942. Physiol. Plant 116: 255-63
25. Sane PV, Ivanov AG, Sveshnikov D, Huner NP, Oquist G. 2002. A transient exchange of the photosystem II reaction center protein D1:1 with D1:2 during low temperature stress of Synechococcus sp. PCC 7942 in the light lowers the redox potential of QB. J. Biol. Chem. 277: 32739-45
26. Sarcina M, Tobin MJ, Mullineaux CW. 2001. Diffusion of phycobilisomes on the thylakoid membranes of the cyanobacterium Synechococcus 7942. Effects of phycobilisome size, temperature, and membrane lipid composition. J. Biol. Chem. 276: 46830-4
27. Sauer J, Gorl M, Forchhammer K. 1999. Nitrogen starvation in Synechococcus PCC 7942: involvement of glutamine synthetase and NtcA in phycobiliprotein degradation and survival. Arch. Microbiol. 172: 247-55
28. Sauer J, Schreiber U, Schmid R, Volker U, Forchhammer K. 2001. Nitrogen starvation-induced chlorosis in Synechococcus PCC 7942. Low-level photosynthesis as a mechanism of long-term survival. Plant Physiol. 126: 233-43
29. Schaefer MR, Golden SS. 1989. Light availability influences the ratio of two forms of D1 in cyanobacterial thylakoids. J. Biol. Chem. 264: 7412-7
30. Schelin J, Lindmark F, Clarke AK. 2002. The clpP multigene family for the ATP-dependent Clp protease in the cyanobacterium Synechococcus. Microbiology 148: 2255-65
31. Schwarz R, Grossman AR. 1998. A response regulator of cyanobacteria integrates diverse environmental signals and is critical for survival under extreme conditions. Proc. Natl. Acad. Sci. U S A 95: 11008-13
32. Shestakov SV, Khyen NT. 1970. Evidence for genetic transformation in blue-green alga Anacystis nidulans R2. Molec. Gen. Genet. 107: 372-5
33. Tchernov D, Helman Y, Keren N, Luz B, Ohad I, et al. 2001. Passive entry of CO2 and its energy-dependent intracellular conversion to HCO3- in cyanobacteria are driven by a photosystem I-generated deltamuH+. J. Biol. Chem. 276: 23450-5
34. Tsinoremas NF, Schaefer MR, Golden SS. 1994. Blue and red light reversibly control psbA expression in the cyanobacterium Synechococcus sp. strain PCC 7942. J. Biol. Chem. 269: 16143-7
35. van Waasbergen LG, Dolganov N, Grossman AR. 2002. nblS, a gene involved in controlling photosynthesis-related gene expression during high light and nutrient stress in Synechococcus elongatus PCC 7942. J. Bacteriol. 184: 2481-90
36. Vazquez-Bermudez MF, Paz-Yepes J, Herrero A, Flores E. 2002. The NtcA-activated amt1 gene encodes a permease required for uptake of low concentrations of ammonium in the cyanobacterium Synechococcus sp. PCC 7942. Microbiol. 148: 861-9
37. Webb R, Troyan T, Sherman D, Sherman LA. 1994. MapA, an iron-regulated, cytoplasmic membrane protein in the cyanobacterium Synechococcus sp. strain PCC7942. J. Bacteriol. 176: 4906-13