Metabolism
1.) BIOSYNTHETIC PATHWAYS IN PHOTOSYNTHETIC BACTERIA (PROKARYOTIC ALGAE):
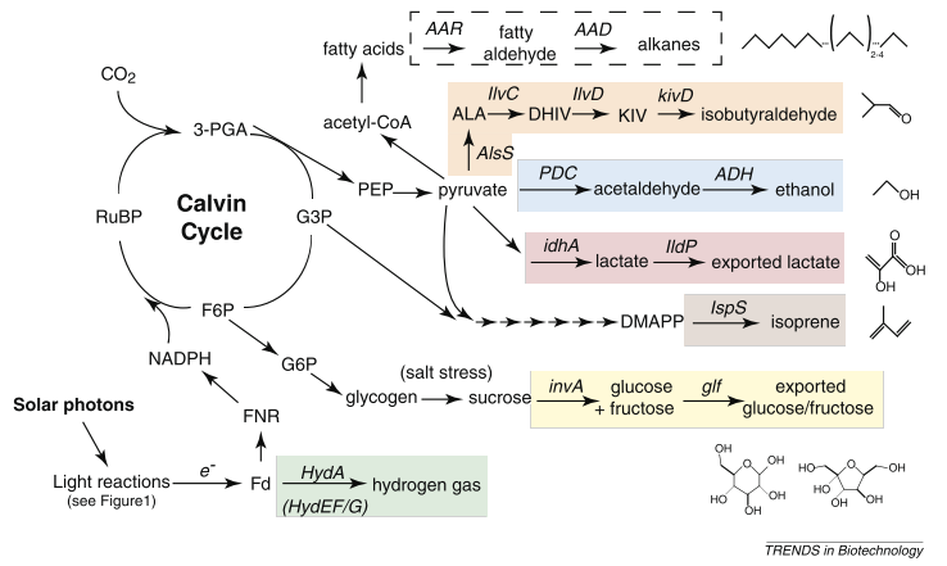
Representation of engineered biochemical pathways in cyanobacteria. Core metabolism of photosynthetic processes is shown in black text. Branch points utilized for the production of various compounds discussed in this review are indicated (highlighted pathways) with relevant enzymes catalyzing specific reactions indicated in italics. Abbreviations: 3-PGA, 3-phosphoglycerate; AAD, aldehyde decarbonylase; ADH, alcohol dehydrogenase II; ALA, 2-acetolactate; AlsS, acetolactate synthase; DHIV, 2,3-dihydroxy-isovalerate; F6P, fructose 6-phosphate; FNR, ferredoxin NADP+ reductase; G6P, glucose 6-phosphate; HydA, [FeFe] hydrogenase; HydEF/G, hydrogenase maturation factors; ldhA, lactate dehydrogenase; IlvD, dihydroxy-acid dehydratase; IlvC, acetohydroxy acid isomeroreductase; PDC, pyruvate decarboxylase; PEP, phosphoenolpyruvate.
Source: Ducat, D. C., Way, J. C., & Silver, P. A. (2011). Engineering cyanobacteria to generate high-value products. Trends in Biotechnology, 29(2), 103–95. doi:10.1016/j.tibtech.2010.12.003
Source: Ducat, D. C., Way, J. C., & Silver, P. A. (2011). Engineering cyanobacteria to generate high-value products. Trends in Biotechnology, 29(2), 103–95. doi:10.1016/j.tibtech.2010.12.003
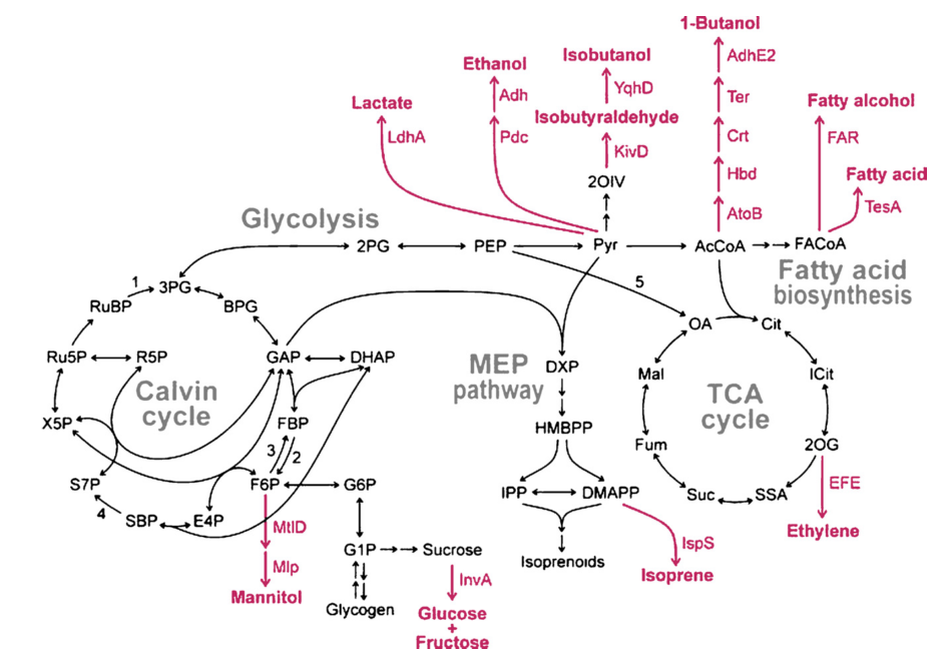
Central metabolic pathways of cyanobacteria. Heterologous pathways and products that have been genetically engineered in cyanobacteria are indicated in magenta. Abbreviations: Enzymes: 1, ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO); 2, fructose-1,6-bisphosphatase; 3, phosphofructok- inase; 4, sedoheptulose-1,7-bisphosphatase; 5, phosphoenolpyruvate carboxylase. Metabolites: 2OG, 2-oxoglutarate; 2OIV, 2-oxoisovalerate; 2PG, 2-phosphoglycerate; 3PG, 3-phosphoglycerate; AcCoA, acetyl-CoA; BPG, 1,3-bisphosphoglycerate; Cit, citrate; DHAP, dihydroxyacetone-phosphate; DMAPP, dimethylallyl-pyrophosphate; DXP, 1-deoxyxylulose-5-phosphate; E4P, erythrose-4-phosphate; F6P, fructose-6-phosphate; FACoA, fatty acyl-CoA; FBP, fructose-1,6-bisphosphate; Fum, fumarate; G1P, glucose-1-phosphate; G6P, glucose-6-phosphate; GAP, glyceraldehyde-3-phosphate; HMBPP, 1-hydroxy-2-methyl-2-butenyl-4-pyrophosphate; ICit, isocitrate; IPP, isopentenyl-pyrophosphate; Mal, malate; OA, oxaloacetate; PEP, phosphoenolpyruvate; Pyr, pyruvate; R5P, ribose-5-phosphate; Ru5P, ribulose-5-phosphate; RuBP, ribulose-1,5-bisphosphate; S7P, sedoheptulose-7-phosphate; SBP, sedoheptulose-1,7-bisphosphate; SSA, succinic semialdehyde; Suc, succinate; X5P, xylulose-5-phosphate.
Source: Rosgaard, L., de Porcellinis, A. J., Jacobsen, J. H., Frigaard, N.-U., & Sakuragi, Y. (2012). Bioengineering of carbon fixation, biofuels, and biochemicals in cyanobacteria and plants. Journal of Biotechnology, 162(1), 134–47. doi:10.1016/j.jbiotec.2012.05.006
Source: Rosgaard, L., de Porcellinis, A. J., Jacobsen, J. H., Frigaard, N.-U., & Sakuragi, Y. (2012). Bioengineering of carbon fixation, biofuels, and biochemicals in cyanobacteria and plants. Journal of Biotechnology, 162(1), 134–47. doi:10.1016/j.jbiotec.2012.05.006
Production of hydrocarbons from cyanobacteria:
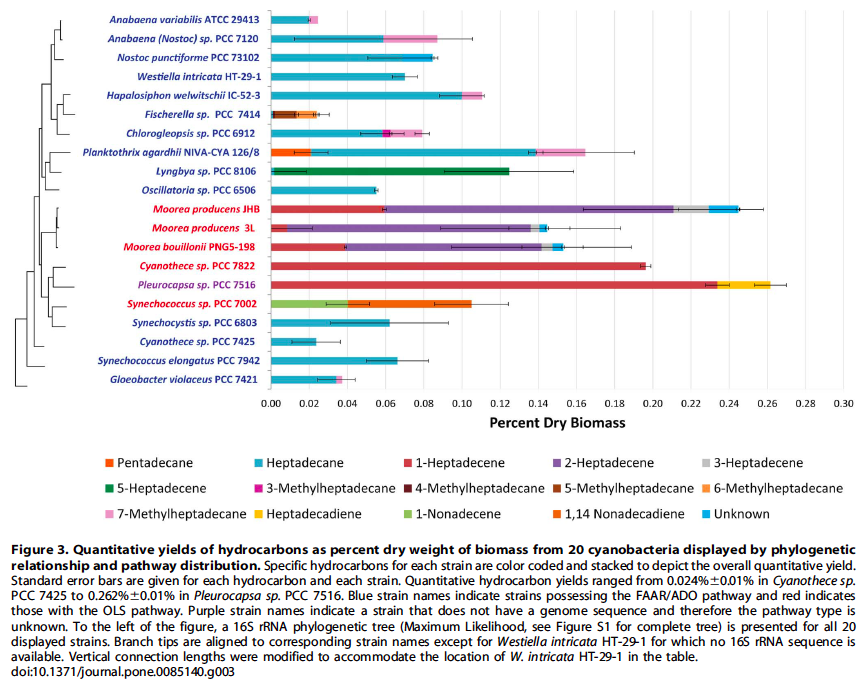
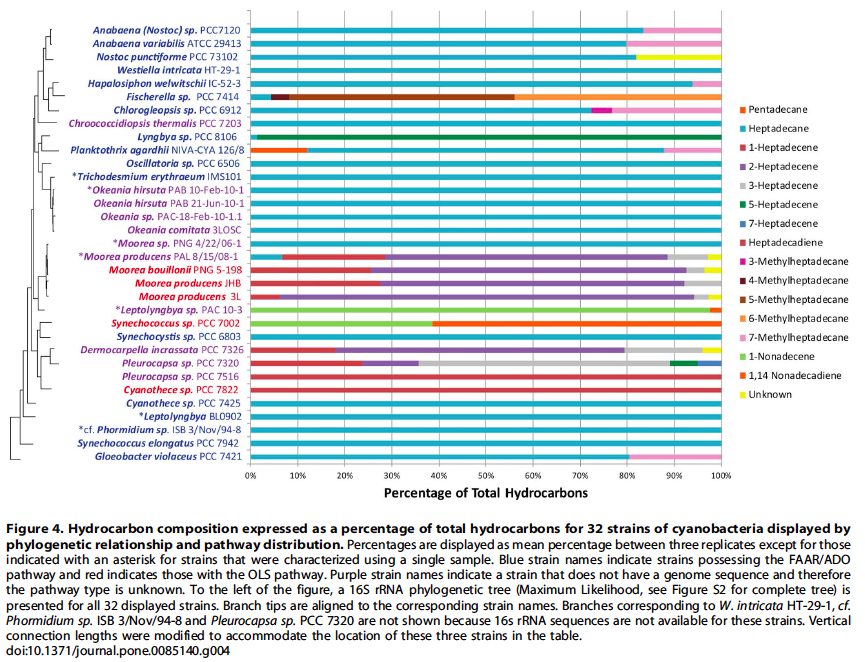
Cyanobacteria possess the unique capacity to naturally produce hydrocarbons from fatty acids. Hydrocarbon compositions of thirty-two strains of cyanobacteria were characterized to reveal novel structural features and insights into hydrocarbon biosynthesis in cyanobacteria. This investigation revealed new double bond (2- and 3-heptadecene) and methyl group positions (3-, 4- and 5-methylheptadecane) for a variety of strains. Additionally, results from a recent study and literature reports indicate that hydrocarbon production is a universal phenomenon in cyanobacteria.
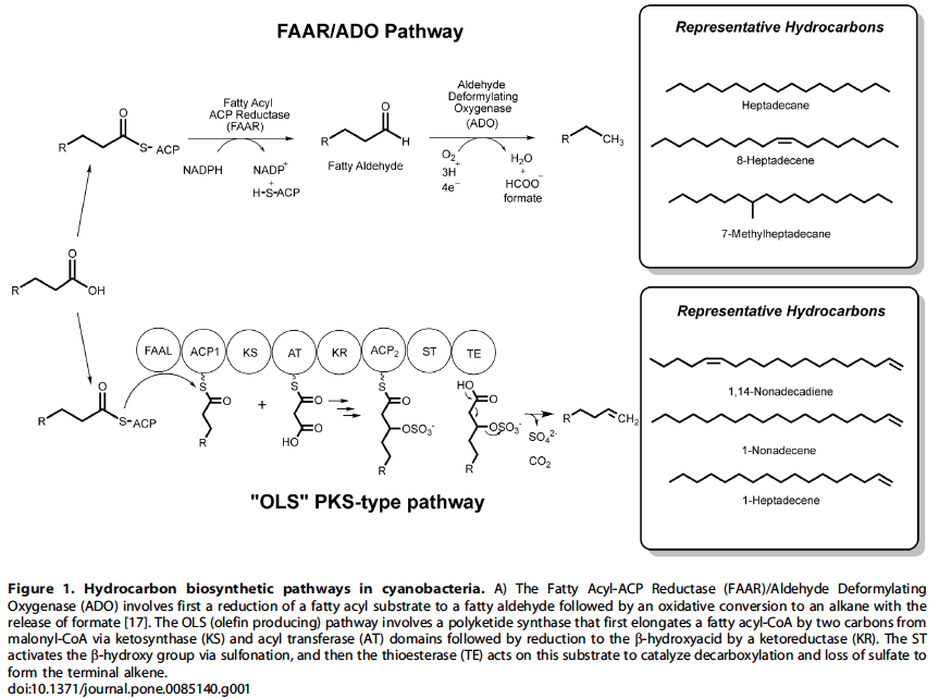
One pathway comprises a two-step conversion of fatty acids first to fatty aldehydes and then alkanes that involves a fatty acyl ACP reductase (FAAR) and aldehyde deformylating oxygenase (ADO). The second involves a polyketide synthase (PKS) pathway that first elongates the acyl chain followed by decarboxylation to produce a terminal alkene (olefin synthase, OLS) - Figure 1.
Hydrocarbon pathways were identified in one-hundred-forty-two strains of cyanobacteria over a broad phylogenetic range (Figure 3 and 4) and there were no instances where both the FAAR/ADO and the OLS pathways were found together in the same genome, suggesting an unknown selective pressure maintains one or the other pathway, but not both.
Source: Coates, R. C., Podell, S., Korobeynikov, A., Lapidus, A., Pevzner, P., Sherman, D. H., … Gerwick, W. H. (2014). Characterization of cyanobacterial hydrocarbon composition and distribution of biosynthetic pathways. PloS One, 9(1), e85140. doi:10.1371/journal.pone.0085140.
2.) BIOSYNTHETIC PATHWAYS IN NUCLEUS CONTAINING ORGANISMS (EUKARYOTIC ALGAE):
2.) BIOSYNTHETIC PATHWAYS IN NUCLEUS CONTAINING ORGANISMS (EUKARYOTIC ALGAE):
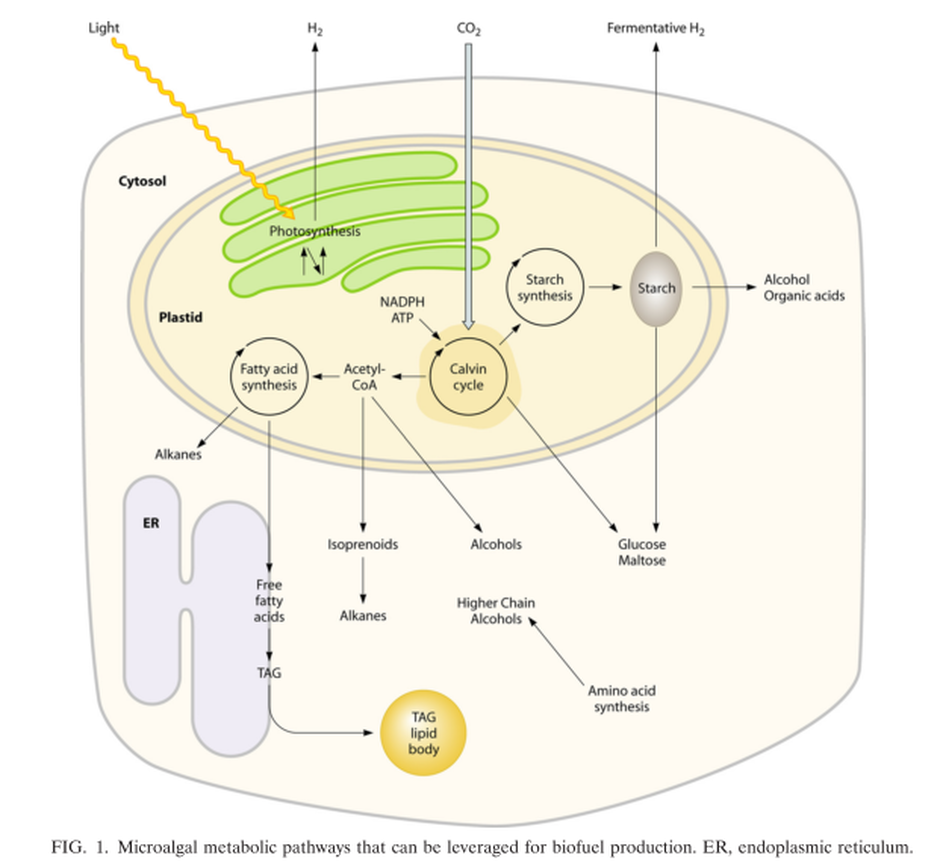
Although the application of genetic engineering to improve energy production phenotypes in eukaryotic microalgae is in its infancy, significant advances in the development of genetic manipulation tools have recently been achieved with microalgal model systems and are being used to manipulate central carbon metabolism in these organisms. It is likely that many of these advances can be extended to industrially relevant organisms.
Source: Radakovits, R., Jinkerson, R. E., Darzins, A., & Posewitz, M. C. (2010). Genetic engineering of algae for enhanced biofuel production. Eukaryotic Cell, 9(4), 486–501. Retrieved from http://ec.asm.org/content/9/4/486.short
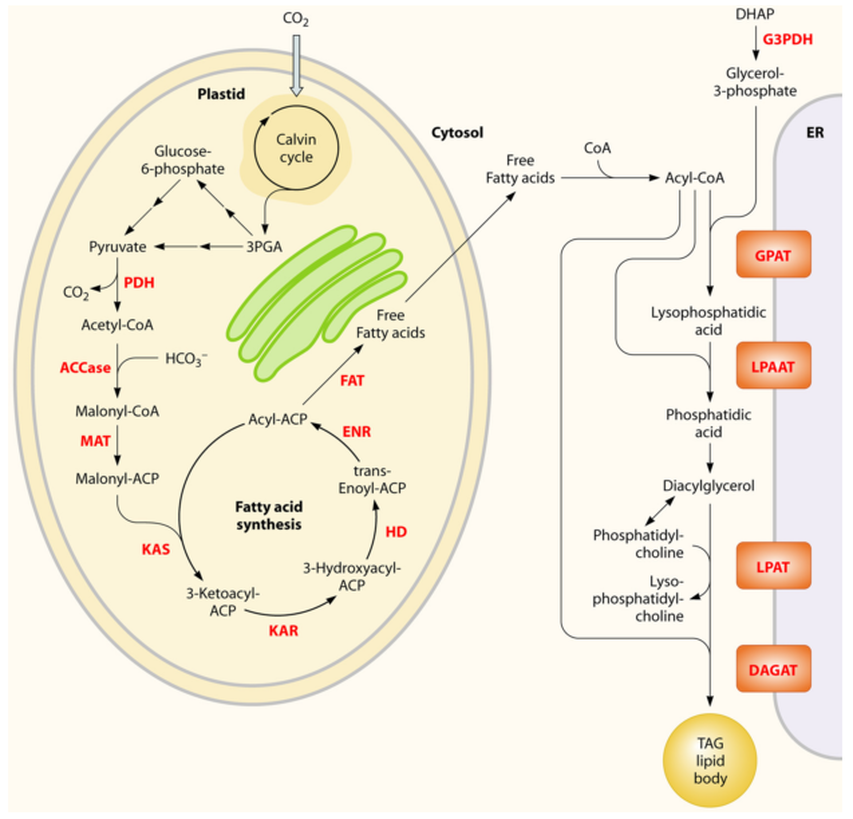
Simplified overview of the metabolites and representative pathways in microalgal lipid biosynthesis shown in black and enzymes shown in red. Free fatty acids are synthesized in the chloroplast, while TAGs may be assembled at the ER. ACCase, acetyl-CoA carboxylase; ACP, acyl carrier protein; CoA, coenzyme A; DAGAT, diacylglycerol acyltransferase; DHAP, dihydroxyacetone phosphate; ENR, enoyl-ACP reductase; FAT, fatty acyl-ACP thioesterase; G3PDH, gycerol-3-phosphate dehydrogenase; GPAT, glycerol-3-phosphate acyltransferase; HD, 3-hydroxyacyl- ACP dehydratase; KAR, 3-ketoacyl-ACP reductase; KAS, 3-ketoacyl-ACP synthase; LPAAT, lyso-phosphatidic acid acyltransferase; LPAT, lyso-phosphatidylcholine acyltransferase; MAT, malonyl-CoA:ACP transacylase; PDH, pyruvate dehydrogenase complex; TAG, triacylglycerols.
During diel light-dark cycles, many microalgae initiate TAG storage during the day and deplete those stores at night to support cellular ATP demands and/or cell division. Consequently, inhibition of beta-oxidation (fatty acid catabolism) would prevent the loss of TAG during the night, but most likely at the cost of reduced growth. This strategy, therefore, may not be beneficial for microalgae grown in outdoor open ponds, but it may be a valid strategy to increase lipid production in microalgae grown in photobioreactors with exogenous carbon sources and/or continuous light.
In some studies, inhibition of lipid oxidation has caused unexpected phenotypes. Several publications have shown that knocking out genes involved in beta-oxidation in S. cerevisiae (baker's yeast) not only can lead to increased amounts of intracellular free fatty acids but also results in extracellular fatty acid secretion in some instances (120, 132, 162). The lipid catabolism genes that have been implicated in fatty acid secretion in S. cerevisiae include acyl-CoA oxidase and several acyl-CoA synthetases.
Source: Radakovits, R., Jinkerson, R. E., Darzins, A., & Posewitz, M. C. (2010). Genetic engineering of algae for enhanced biofuel production. Eukaryotic Cell, 9(4), 486–501. Retrieved from http://ec.asm.org/content/9/4/486.short
During diel light-dark cycles, many microalgae initiate TAG storage during the day and deplete those stores at night to support cellular ATP demands and/or cell division. Consequently, inhibition of beta-oxidation (fatty acid catabolism) would prevent the loss of TAG during the night, but most likely at the cost of reduced growth. This strategy, therefore, may not be beneficial for microalgae grown in outdoor open ponds, but it may be a valid strategy to increase lipid production in microalgae grown in photobioreactors with exogenous carbon sources and/or continuous light.
In some studies, inhibition of lipid oxidation has caused unexpected phenotypes. Several publications have shown that knocking out genes involved in beta-oxidation in S. cerevisiae (baker's yeast) not only can lead to increased amounts of intracellular free fatty acids but also results in extracellular fatty acid secretion in some instances (120, 132, 162). The lipid catabolism genes that have been implicated in fatty acid secretion in S. cerevisiae include acyl-CoA oxidase and several acyl-CoA synthetases.
Source: Radakovits, R., Jinkerson, R. E., Darzins, A., & Posewitz, M. C. (2010). Genetic engineering of algae for enhanced biofuel production. Eukaryotic Cell, 9(4), 486–501. Retrieved from http://ec.asm.org/content/9/4/486.short